Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Darwiniana, nueva serie
Print version ISSN 0011-6793On-line version ISSN 1850-1699
Darwiniana, nueva serie vol.4 no.2 San Isidro Dec. 2016
ECOLOGÍA Y FITOGEOGRAFÍA
Local hibridization in subtropical mountain habitats: can cedrela (meliaceae) maintain species’ identity in sympatry?
M. Paula Quiroga1, Andrea C. Premoli2, Alfredo Grau3 & Lucio Malizia4
1 Instituto de Investigaciones en Biodiversidad y Medio Ambiente (INIBIOMA-CONICET) Laboratorio Ecotono, Departamento de Botánica, Centro Regional Universitario Bariloche, Universidad Nacional del Comahue, Quintral 1250, 8400 Bariloche, Río Negro, Argentina; emepequ@gmail.com (author for correspondence).
2 Instituto de Investigaciones en Biodiversidad y Medio Ambiente (INIBIOMA-CONICET) Laboratorio Ecotono. Departamento de Biología, Centro Regional Universitario Bariloche, Universidad Nacional del Comahue. Quintral 1250, 8400 Bariloche, Río Negro, Argentina.
3 Instituto de Ecología Regional, Facultad de Ciencias Naturales, Universidad Nacional de Tucumán, Miguel Lillo 205, 4000 San Miguel de Tucumán, Tucumán, Argentina.
4 Centro de Estudios Territoriales Ambientales y Sociales, Facultad de Ciencias Agrarias, Universidad Nacional de Jujuy, Alberdi 47, Y4600DTA San Salvador de Jujuy, Jujuy, Argentina.
Abstract
Congener species with incomplete reproductive barriers that coexist along environmental gradients may be prone to ecological divergence, in spite of the potential for hybridization in sympatry. We analyzed distribution patterns of isozymes, plastid, and nuclear DNA sequences at regional and local scales in three timber Cedrela species of the subtropics in the northern Argentina (Cedrela angustifolia, C. balansae, and C. saltensis), to test whether populations of distinct species have diverged in montane habitats in relation to their ecological characteristics. Cedrela balansae and C. angustifolia can be identified by diagnostic isozyme alleles. Nuclear ITS sequences yielded intraindividual polymorphism; ambiguous bases were shared between C. balansae and C. saltensis while those of C. angustifolia were part of its intraspecific polymorphism. Chloroplast DNA consisted mainly of a low-elevation haplotype present in C. balansae and C. saltensis and other found in C. angustifolia which in turn was shared locally by all species in sympatry. Multivariate UPGMA analysis of isozymes and Bayesian phylogeny of haplotype ITS yielded concordant patterns. Populations of C. angustifolia clustered in one group and were separated from the rest whereas those of C. saltensis and C. balansae were grouped together in one cluster. This indicates that stronger reproductive barriers exist between C. angustifolia and the low-elevation taxa C. saltensis and C. balansae which seem to maintain continuous gene flow. Nonetheless, under particular environmental settings, i.e., the three species hybridized sometime in the past and later became differentiated through ecological divergence.
Keyword. Argentina; Cedrela angustifolia; Cedrela balansae; Cedrela saltensis; Yungas.
Hibridación local en hábitats de montaña subtropicales: ¿Cedrela (Meliaceae) puede mantener la identidad de las especies en simpatría?
Resumen
Las especies congéneres con barreras reproductivas incompletas que coexisten a lo largo de un gradiente ambiental pueden ser proclives a la divergencia ecológica, a pesar del potencial de hibridación en simpatría. Estudiamos los patrones de distribución de isoenzimas y de secuencias de ADN nuclear y plastidial a escala local y regional en las tres especies de árboles del género Cedrela del subtrópico de noroeste de Argentina para analizar si las poblaciones de las distintas especies han divergido en sus hábitats montanos en relación a sus características ecológicas. Cedrela balansae y C. angustifolia pueden ser identificadas por alelos isoenzimáticos diagnósticos. Las secuencias nucleares del ITS mostraron polimorfismo intraindividual; bases ambiguas fueron compartidas entre C. balansae y C. saltensis, mientras que en C. angustifolia es parte de su polimorfismo intraespecífico. El ADN del cloroplasto consistió en un haplotipo compartido entre las especies de baja altitud C. balansae y C. saltensis y otro diferente para C. angustifolia el cual comparte con las otras dos especies cuando están en simpatría. Los análisis de UPGMA de isoenzimas y filogenético bayesiano de haplotipos del ITS muestran patrones concordantes. Las poblaciones de C. angustifolia se agruparon dentro de un mismo cluster y se separaron de las de C. saltensis y C. balansae que se mantuvieron agrupadas. Esto indica que existen barreras reproductivas entre C. angustifolia y los otros taxa de baja elevación C. saltensis y C. balansae, las cuales mantienen un continuo flujo génico. No obstante, bajo los distintos ambientes que ocupan, las tres especies hibridaron en algún momento en el pasado y más tarde comenzaron a diferenciarse a través de divergencia ecológica.
Palabras clave: Argentina; Cedrela angustifolia; Cedrela balansae; Cedrela saltensis; Yungas.
Original recibido el 8 de junio de 2016,
aceptado el 14 de octubre de 2016
Editora Asociada: Lone Aagesen
INTRODUCTION
Classical debates on speciation focused on the geography of speciation in relation to reproductive isolation, and thus drift, causing divergence (Mayr, 1947). While allopatric speciation mostly takes place when physical barriers impede genetic exchange, and therefore prompt divergence, sympatric speciation occurs in the face of gene flow. Under the absence of reproductive isolating mechanisms, divergence may result from the action of ecologically-based divergent selection which arises as a consequence of individuals interacting with their environment, i.e. ecological speciation (Orr & Smith, 1998). Ecological selection is divergent when it acts in contrasting environmental settings, due to abiotic factors e.g. climate and distribution of resources as well as biotic effects such an individual’s ability to obtain food and/or nutrients, attract pollinators, or avoid predators and various forms of interspecific interactions (Schluter, 2001). This differs from other models of speciation in which the evolution of reproductive isolation involves key processes other than ecological selection, including speciation by polyploidization, hybridization, genetic drift, and founder events/population bottlenecks (Rundle & Nosil, 2005).
Sympatric/parapatric congener species inhabiting mountain habitats provide the opportunity to study isolation mechanisms as a result of ecological segregation of niches in relation to distinct environmental factors that abruptly change with elevation. Nonetheless, if such taxa maintain incomplete reproductive barriers potential hybridization events may occur wherever they overlap. The occurrence of intermediate forms between sympatric species, i.e. potential hybrids, either result from the cross of taxa undergoing the process of splitting by sympatric speciation and thus show incipient reproductive isolation or alternatively, they are the product of secondary contact between taxa that have diverged sometime in the past but have not developed complete reproductive barriers (Coyne & Orr, 2004). Cedrela species are taxonomically challenging because they can be easily confounded morphologically (see key to the species of Cedrela in Pennington& Muellner, 2010). Therefore, a concern arose about the possibility of hybridization between species with nearly similar ecological tolerances.
Recent studies in Cedrela suggest that interspecific reproductive isolation seems to be incomplete, particularly among some of the South American species (Köcke, 2015). Furthermore, neither species’ monophyly of e.g. C. fissilis, nor the delimitation of species such as C. odorata and C. fissilis was supported by phylogenetic analyses (Garcia-Gonçalvez et al., 2011). In addition, cryptic speciation, i.e. the existence of genetically distinct but morphologically indistinguishable entities, was suggested for C. odorata (Muellner et al., 2009, 2010; Cavers et al., 2013) and C. fissilis (Garcia-Gonçalvez et al., 2011). Furthermore, Cedrela species can tolerate a wide array of environmental variables, i.e. precipitation and temperature (Koecke et al., 2013) which may favor range overlap and thus the potential formation of intermediate forms.
Three species of Cedrela inhabit subtropical forests of the southern Yungas in northwestern Argentina: C. balansae, C. angustifolia (following Pennington & Muellner, 2010), and C. saltensis (Zapater et al., 2004). These are among the most valuable subtropical hardwoods of Argentina and therefore, they have been heavily logged (Malizia et al., 2006). They have a variable degree of geographic overlap along latitudinal and elevation gradients: C. angustifolia is found in Montane forests between 500 and 2300 m above sea level (a.s.l.) reaching its southern-most distributional range at 28º15’ S; C. balansae occurs in Pedemont forests between 400 and 1100 m a.s.l. attaining its southern limit at 24º 30’ S; and C. saltensis occupies the transition of Montane and Pedemont forests between 700 and 1100 m a.s.l. with its southern limit at 24º 40’ S (Fig. 1; Grau et al., 2006). The latter has a restricted range (<1000 km2) most of which occurs in sympatry (600 km2) with the other two species (Grau et al., 2006). In comparison with other Cedrela from South and Central America, the three species have similarly high variability in their climatic envelopes, e.g. temperature seasonality (Koecke et al., 2013). In a previous study, we analyzed the possible hybrid origin of Cedrela saltensis from C. balansae and C. angustifolia, based on a reduced number of populations and using isozymes. This preliminary analysis yielded the presence of diagnostic alleles in C. saltensis that suggested that this species is taxonomically distinct (Premoli et al., 2011). This result was in agreement with a phylogenetic analysis showing that C. saltensis probably had a separate evolutionary history (Muellner et al., 2009).
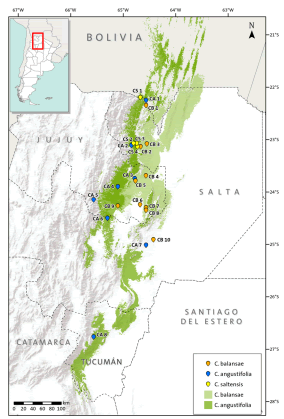
Fig 1. Distribution ranges of Cedrela angustifolia, C. balansae, and C. saltensis, and locations of the studied populations. The numbers correspond to populations indicated in Table 1. Color version at http://www.ojs.darwin.edu.ar/index.php/darwiniana/article/view/715/700
We analyzed distribution patterns of molecular diversity along the range of each of the three pure Cedrela species (i.e. at regional scale) and at a local scale where at least two such species coexist in sympatry. We test the hypothesis whether populations of C. saltensis, C. balansae, and C. angustifolia which in turn coexist with a variable degree of range overlap have diverged in montane habitats. We aim to solve the following questions: 1) Can the three Cedrela species be distinguished genetically by neutral markers in relation to the different habitats they occupy (i.e. elevation)? 2) Do they maintain gene flow particularly where their ranges overlap? 3) If so, does gene flow occur at present and/or has also occurred in the past? We used isozymes, plastid, and nuclear DNA sequences of samples collected within the three species’ natural ranges to answer these questions.
MATERIALS AND METHODS
Sample collection
To study the distribution of genetic diversity of Cedrela angustifolia, C. balansae, and C. saltensis, we sampled what is considered putative pure species inhabiting seasonally dry forests of northern Argentina. They were identified by morphological characters and altitudinal distribution. During years 2008-2009 we collected fresh leaf tissue from several populations at randomly selected sites of each species (Table 1), collecting a total of 227 individuals. At each location all species present were sampled.
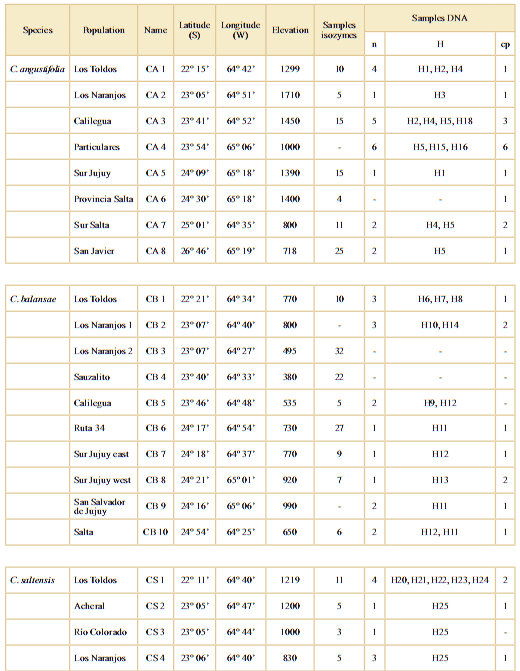
Table 1. Location of sampled populations of Cedrela angustifolia (CA), C. balansae (CB), and C. saltensis (CS) in northwestern Argentina. n: nuclear; H: haplotypes to ITS region; cp: chloroplast. Accession number H1 to H25: KX840898 to KX840922.
Isozymes
Protein extraction usually took place within a week of field work. However, because of the deciduous habit leaflets were released of the rachis before protein extraction, resulting in the loss of enzyme activity of some samples. Enzyme extracts were prepared by grinding ca. 1g of leaf tissue using 1–1.5 mL of extraction buffer (Mitton et al., 1979). Homogenates were stored at -80ºC until they were absorbed onto Whatman No. 3 paper wicks that were loaded into 12% starch gels (S- 5651 starch potato; Sigma, St. Louis, MO, USA). Five enzymes were resolved in two combinations of buffer systems that yielded information on a total of eight isozyme loci. These were Isocitrate dehydrogenase (Idh1, Idh2), Malic enzyme (Me1, Me2), and Shikimate dehydrogenase (Skdh) that were resolved in a buffer morpholine-citric system pH 7.5 (Ranker et al., 1989), which was run at a constant current of 25 mA for about 6 h. Peroxidase (Per1, Per2) and Phosphoglucoisomerase (Pgi) in buffer system pH 8.2 (Poulik, 1957) was run at a constant current of 20 mA for about 6 h.
Differences among populations and species were analyzed by heterogeneity of allelic frequencies by chi-square tests. Levels of within-population isozyme variation and at the species level were described by standard population genetic measures using Popgene v. 1.32 (Yeh & Boyle, 1997). These were the mean number of alleles (A), the effective number of alleles (NE), the proportion of polymorphic loci using the sensu stricto criterion (PSS), and the observed and expected heterozygosities under Hardy–Weinberg equilibrium conditions (HO and HE, respectively). Genetic diversity indices were compared statistically among species, which first calculates the average for each species over samples and loci calculated from the randomized data set with 9999 permutations using FSTAT v. 2.9.3.2 (Goudet, 1995).
The degree of population divergence (FST; Wright, 1965) between all population pairs was estimated using polymorphic loci by FSTAT v. 2.9.3.2 (Goudet, 1995). We compared average within- and between-species FST values by Kruskal Wallis non-parametric tests with pairwise multiple comparisons of mean rank sums. At the local level, we tested the degree of genetic divergence by paired between-species FST at the two localities were all species coexist in sympatry, i.e. Los Toldos (CA1, CB1, CS1) and Los Naranjos (CA2, CB3, CS4) populations (at 22 and 23º South latitude, respectively; Table 1).
Genetic relatedness among populations of the three species was analyzed with multivariate UPGMA cluster analysis using modified Rogers genetic distance (Wright, 1978).
DNA isolation, amplification and sequencing
We performed DNA extractions following the ATMAB method (Dumolin et al., 1995). Non-coding regions of chloroplast DNA (cpDNA) were amplified by polymerase chain reaction (PCR) using four intergenic spacer regions, namely: psbB-psbH (BH; Hamilton, 1999), psbA-trnH (AH; Hamilton, 1999), trnD-trnT (DT; Demesure et al., 1995), and trnL-trnF (LF; Taberlet et al., 1991). Nuclear DNA (nDNA) was amplified for the internal transcribed spacer 1, 5.8S ribosomal RNA gene, and internal transcribed spacer 2, complete sequence (ITS; Muellner et al., 2005). The PCR reaction mix contained 1 mL of DNA extract (~10 ng), 2.5 ml buffer (Invitrogen) 10x, 2.5 ml dNTPs 10x, 1.2 ml MgCl2 50 mmol, (0.75 mM of each primer 10mM), and 0.1 ml of Taq DNA polymerase (Invitrogen), as well as 2 ml dimethyl sulfoxide (DMSO) only to analyze the ITS region, on a total volume of 25 mL. The PCR amplification conditions were: 4 min at 95 ºC, 35X (1 min at 94 ºC; 1 min annealing at 53 ºC for BH, AH, DT, LF and 49ºC for ITS; 1.5 min at 72 ºC), and 10 min at 72 ºC. The PCR products were purified by a reaction using 2.5 U of Exonuclease I (USB) and (0.25 U of Shrimp Alkaline Phosphatase (USB) for 10 mL of PCR product (conditions: 15 min at 37 ºC; 15 min at 85 ºC). The amplified DNA was sequenced with an ABI PRISM 3100 Avant Genetic Analyzer (Applied Biosystems, Laboratorio Ecotono, Universidad Nacional del Comahue, Bariloche) using a BigDye Terminator v3.1 Cycle sequencing kit (Applied Biosystems). The cycle sequencing reactions were performed following the manufacturer’s protocols. Sequences of each sampled haplotype were deposited in GenBank (Accession number KX840881 to KX840927).
Sequence data were aligned with Mega4 (Tamu ra et al., 2007) and concatenated manually into a single combined dataset for posterior analyses only for cpDNA regions for each species. Chloroplast and genomic DNA haplotypes were determined from both nucleotide substitutions and indels. We calculated different diversity parameters including variation in alignment size, number of haplotypes (H), number of variable sites (V) and parsimony informative sites (PI), and nucleotide (π) and haplotype (h) diversity with DNAsp 5.10 (Librado & Rozas, 2009) for each species.
Phylogenetic analysis
We inferred the phylogenetic relationships among haplotypes of C. balansae, C. angustifolia, and C. saltensis by Bayesian inference (BI) analysis using MrBayes v.3.1.2 (Ronquist & Huelsenbeck, 2003). This analysis was carried out independently for the cpDNA and ITS datasets.
The alignment analyses of cpDNA yielded a total of 3050 bp but only 4 parsimony-informative characters. The most appropriate models of nucleotide substitution identified by Modeltest v3.7 (Posada and Crandall, 1998) and MrModeltest v2.3 (Nylander, 2004) under the Akaike Information Criterion (Akaike, 1973) (AIC) was implemented in the BI analyses. The AIC indicated that the best-fit model was HKY+I among the 24 models of molecular evolution. The program MrBayes v3.1.2 was used to estimate an unrooted Bayesian phylogeny using the model selected. Bayesian analysis was performed using two simultaneous runs of 2 million generations each, with one cold and three heated chains in each run. Sampling was carried out once every 100 trees; the first 2500 trees were discarded as burn-in samples. A 50% majority-rule consensus tree of the two independent runs was obtained with posterior probabilities that were equal to bipartition frequencies.
We used sequences of the ITS region of the three species to perform BI at the regional level. The outgroup consisted of sequences of the closely related genus Toona (accession number: FJ518906, FJ462489, FJ462488), C. salvadorensis, and C. dugesii (accession numbers FJ462484 and FJ462483, respectively). The BI analysis was performed using four replicates with four chains (three heated and one cold) that were run for seven million generations sampling every 100 generations with a burn-in period of approximately 10% of the trees for the ITS region. Nodal support was determined using Bayesian posterior probabilities as implemented by MrBayes. The most appropriate models of nucleotide substitution were identified with the same methodology used for cpDNA. The best-fit model of AIC was HKY+I+G for data including the ambiguous bases and hybrid individuals and the model HKY+I without ambiguous bases and hybrid individual. A 50% majority-rule consensus tree of the two independent runs was obtained with posterior probabilities that were equal to bipartition frequencies. At the local scale, we preformed analysis of BI (using the same conditions as described above) including all individuals from the Los Toldos population which is the location where the three species coexist in sympatry and where ambiguities, i.e. intraindividual variation at single bases were detected. This ambiguity was solved following Garcia-Gonçalvez et al. (2011) so that each of the five sequences that harbored double peaks at a given location was separated into a pair of new sequences. These sequences were identical to the respective parental sequence except that each sequence of the pair included one of the two possible nucleotide codes instead of the formerly ambiguous code (Garcia-Gonçalvez et al., 2011).
RESULTS
Isozyme variation
To obtain reliable isozyme profiles, several individuals per population and species were analyzed, although that enzyme activity depended on the freshness of samples upon arrival to the laboratory. Isozyme profiles yielded diagnostic alleles, i.e. those present exclusively in just one species, Idh1-1in three populations of C. angustifolia and Idh2-1, Me2-1, and Per1-2 which in turn were unique to just one population of C. balansae. Neither diagnos- tic (i.e. species-specific) nor unique (i.e. population-specific for a given population within a species) alleles were present in the range-restricted C. saltensis [see Supporting Information]. Allelic frequencies at isozyme loci differed among populations of the three analyzed Cedrela species. Seven of eight tests of heterogeneity in allelic frequencies yielded significant differences among populations by means of chi-square tests [see Appendix].
All three species were genetically diverse by isozymes (Table 2). Relatively reduced among-population divergence was measured within each species: C. angustifolia FST = 0.170 [CI 0.057-0.292], C. balansae FST = 0.115 [CI 0.040-0.232], and C. saltensis FST = 0.255 [CI 0.059-0.444] (Fig. 2).
Table 2. Parameters of genetic variation within populations and species of Cedrela from the southern Yungas, Argentina. These are: mean number of alleles (A), effective number of alleles (NE), percentage of polymorphic loci sensus tricto (Pss), and observed and expected heterozygosity (HO and HE, respectively). Population averages and species values are shown at the bottom of each column. Standard errors are shown in parentheses. Cedrella angustifolia (CA), C. balansae (CB) and C. saltensis (CS). Location of the populations are shown in Table 1.
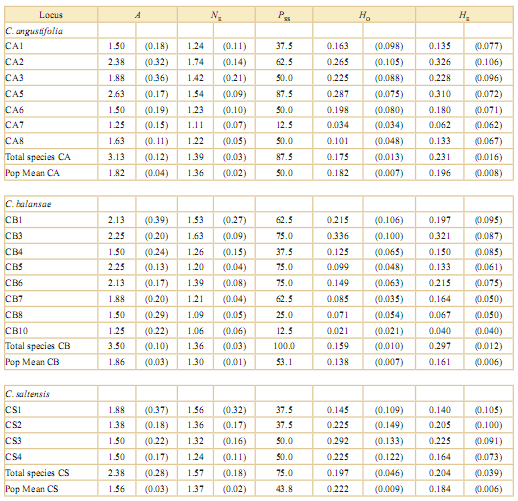
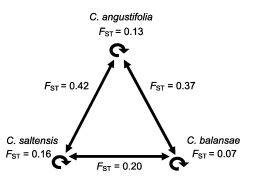
Fig. 2. Diagram of population divergence by (FST) within- (circular arrows) and between-species (linear arrows) of Cedrela.
Statistically lower between-species divergence (p<0.001, Kruskal-Wallis test: H(2, N=113) =33.66) was obtained for C. balansae and C. saltensis population pairs (FST = 0.20) than between C. angustifolia and each of the two former species (C. angustifolia v C. balansae FST = 0.37 and C. angustifolia v C. saltensis FST = 0.42). A similar pattern was detected at the local level with greater divergence between C. angustifolia and any of the other species growing in sympatry (C. angustifolia v C. balansae FST = 0.32 and 0.46, C. angustifolia v C. saltensis FST = 0.45 and 0.40) than between C. balansae and C. saltensis FST = -0.1 and 0.11 at Los Toldos and Los Naranjos, respectively.
Multivariate cluster analysis (UPGMA) using modified Rogers genetic distance showed that populations of C. angustifolia cluster together in one group which resulted isoenzimatically distinct from populations of C. saltensis and C. balansae (Fig. 3).
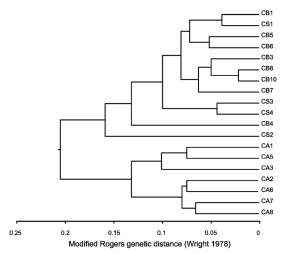
Fig. 3. Multivariate cluster analysis (UPGMA) by means of modified Rogers distance (Wright, 1978) of C. angustifolia (CA), C. balansae (CB), and C. saltensis (CS) populations. Numbers correspond to location of sampled populations as described in Table 1.
DNA variation
Sequences of cpDNA yielded low polymorphism and reduced intra- and inter-specific variation (Table 3, Fig. 4). The final cpDNA dataset that resulted from the concatenation of the four chloroplast regions was 3045 bases long and included four haplotypes (Table 3). Nonetheless, chloroplast haplotypes were geographically structured two of them were widespread (H1 and H2) and the other two were unique to population CB2 of C. balansae and CA3 of C. angustifolia (H3 and H4, respectively). Wide ranging H1 was mostly exclusive to populations of C. angustifolia except at the high-elevation northern-most studied populations at Los Toldos where the three species coexist in sympatry/parapatry and share this chloroplast haplotype. The other H2 widespread haplotype was exclusively present in C. balansae and C. saltensis wherever they occur as pure populations (Fig. 4).
Table 3. Genetic diversity of cpDNA for three species of Cedrela. N: Number of haplotypes; # VS/ PI: # Variable sites / Parsimony informative sites; h: Haplotype diversity; p: Nucleotide diversity. Standard errors are shown in parenthesis.

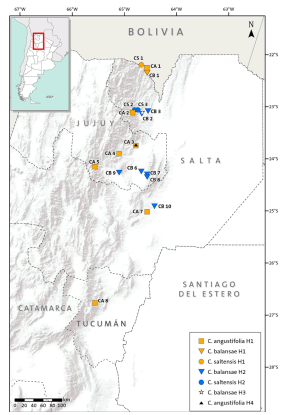
Fig. 4. Geographical distribution of chloroplast DNA haplotypes. The numbers correspond to populations indicated in Table 1. Haplotypes (H) correspond to Table 3. Color version at http://www.ojs.darwin.edu.ar/index.php/darwiniana/article/view/715/700
The aligned nITS region matrix consisted of 578 characters. Forty-seven sites were variable, 26 of which were potentially parsimony informative. In total, 25 haplotypes were identified (Table 4). Out of the total variable sites, 43% were shared between the three species, 27% between C. balansae and C. saltensis, 15% between C. angustifolia and C. balansae, 10% between C. angustifolia and C. saltensis, whereas 5% differed among the three species. Bayesian consensus trees using the nITS dataset showed that haplotypes of C. angustifolia constitute a monophyletic group while haplotypes of C. balansae and C. saltensis constitute a shared group (Fig. 5).
Table 4. Genetic diversity of the ITS1-2 region for three species of Cedrela. N: number of haplotypes; # VS/ PI: Number of variable sites / Parsimony informative sites; h: Haplotype diversity; p: Nucleotide diversity. Standard errors are shown in parenthesis.
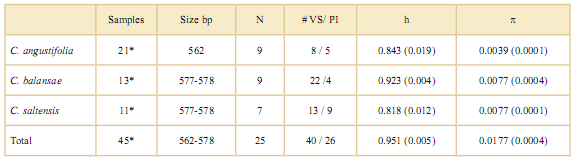
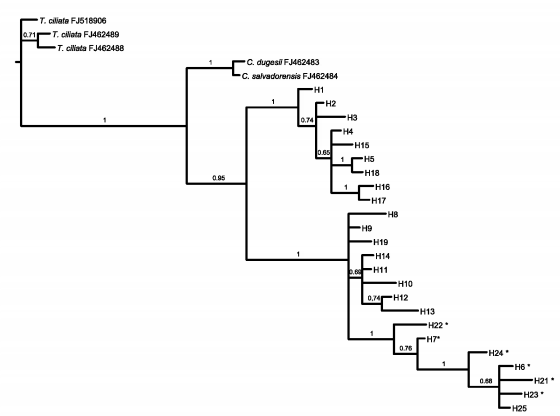
Fig. 5. Bayesian phylogeny (consensus tree) of nITS data of C. angustifolia, C. balansae, and C. saltensis, including Toona ciliata (3 samples), C. salvadorensis, and C. dugesii as outgroup. Numbers correspond to locations of the sampled populations following Table 1. Numbers above branches indicate posterior probabilities values of Bayesian analyses. * correspond to individuals with ambiguities.
Seven of the 47 variable sites showed at least one ambiguity, that is, they presented overlapping double peaks at the same site. The pattern of double peaks in the electropherograms suggested intraindividual polymorphism. We detected double peaks (ambiguities) in different individuals of the three species. Ambiguities of C. saltensis and C. balansae consist of double peaks present in each of the pure species (interspecific variation), whereas those of C. angustifolia corresponded to the within-species variation (intraspecific variation).
At Los Toldos, out of a total of 578bp of the ITS region, 25bp were variable (Table 5). In 10 of those sites one individual of C. balansae presented ambiguities, i.e. double peaks that were always shared with bases of C. saltensis. Also, two individuals of C. saltensis presented ambiguities at four sites which in turn were shared with C. balansae. In contrast, one individual of C. angustifolia from Los Toldos present ambiguities that corresponded to intraspecific variation. Nonetheless, the phy- logenetic relationships among individuals of the three species at Los Toldos by ITS showed that C. angustifolia individuals are different from those of the other two species (Fig. 6).
Table 5. Samples of nuclear ITS sequences with nonambiguous and ambiguous bases of three Cedrela species’ populations. Sequences with ambiguities labeled as A or B are indicated in boxes. The location of the populations are shown in Table 1. The number after the hyphen corresponds to distinct individuals.
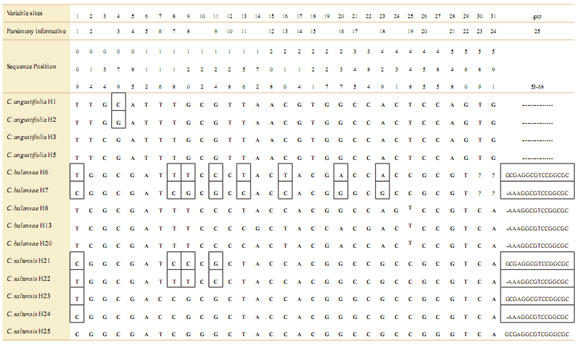
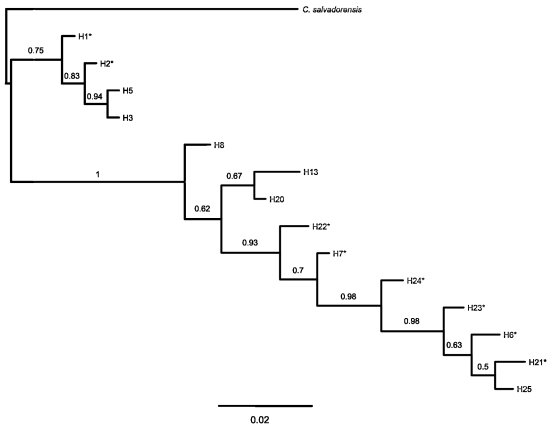
Fig. 6. Bayesian phylogeny (consensus tree) of nITS data of sympatric populations of C. angustifolia, C. balansae, and C. saltensis from Los Toldos population, allopatric populations of each species, and C. salvadorensis as outgroup. Numbers correspond to location of sampled populations following Table 1. * correspond haplotypes with ambiguous bases, i.e. alternative possible nucleotide sequences. Numbers above branches indicate posterior probabilities values of Bayesian analyses.
DISCUSSION AND CONCLUSIONS
Concordant genetic relationships were depicted by ITS and isozymes and show that high-elevation C. angustifolia has diverged from the low-elevation C. balansae and C. saltensis, which in turn seem to be genetically more similar. This result is supported by phylogenetic analysis of ITS haplotypes and clustering schedule of populations using isozyme data (Fig. 2, 3 and 5). At the local scale in Los Toldos and Los Naranjos, where the three taxa coexist in sympatry the same pattern was obtained; i.e. a greater divergence measured by FST values between C. angustifolia and each of the other two species than between C. balansae and C. saltensis. These results suggest the relevance of distinct environmental settings, i.e. elevation, as promotor of ecological divergence in Cedrela. The distribution of cpDNA haplotypes indicated a shared evolutionary history among the species. In particular, concordant patterns of the widely distributed H2 haplotype in C. balansae and C. saltensis provide evidence of potential interspecific gene flow at the regional level. In addition, while the widespread H1 haplotype is mostly restricted to C. angustifolia, it was shared by all species in the northern-most population suggesting that hybridization has probably also occurred locally among the three species (see below).
High levels of genetic diversity by means of isozymes and nuclear DNA correlate with the elevated variability suggested for the three studied species in relation to tolerance to a wide range of temperatures, e.g. temperature seasonality as the difference between temperature maximum of the warmest month and temperature minimum of the coldest month (Koecke et al., 2013). Nonetheless, they generally occupy spatially segregated microhabitats at three elevation strata of the southern Yungas: C. angustifolia at higher elevations while C. saltensis are more common at middle and lower elevations and C. balansae at the lowest end. Quantitative traits measured under common gardens on seedlings of C. odorata from distinct provenances, mesic and dry, suggest adaptive genetically-based differences which were interpreted as potential incipient speciation in relation to contrasting habitats (Navarro et al., 2002).
The nuclear ITS region yielded intraindividual polymorphism, e.g. ambiguous bases, as previously reported in Cedrela (Garcia-Gonçalvez et al., 2011; Zelener et al., 2016). Interestingly, the three species show ambiguous bases at Los Toldos where they occur in sympatry. While such intraindividual polymorphism was shared between C. balansae and C. saltensis and thus suggesting local interspecific gene flow, that of C. angustifolia was part of intraspecific polymorphism within this species’ gene pool. The intraspecific ambiguities detected in C. angustifolia are similar to those found in other Cedrela species such as C. fissilis (Garcia-Gonçalvez et al., 2011). These authors argue that hybridization, polyploidy, and long generation times in plants are factors that contribute to incomplete concerted evolution that has probably contributed to the intra-individual polymorphism in this species. Other Cedrela species analyzed by Muellner et al. (2009) presented ambiguities (C. angustifolia FJ462478, C. balansae FJ462474, C. odorata FJ462464), reflecting high variation in ITS sequences within the genus. Hence, both interspecific and intraspecific hybridization bring together different alleles from the parental sources into a single genome and therefore can give rise to intraindividual polymorphism (Koch et al., 2003; Muir et al., 2001), as in our case.
The three species at Los Toldos shared the same chloroplast haplotype at this location which indicates that either hybridization and thus chloroplast capture probably occurred sometime in the past among the three species and/or that they derived relatively recent from a common ancestor and thus they presents incomplete lineage sorting. The genetic similarity by isozymes and shared interspecific polymorphism in ITS sequences between C. balansae and C. saltensis are in accordance with Koenen et al. (2015) and support the hypothesis of their relatively recent origin product of a contemporaneous expansion possibly during the Holocene. On the other hand, the greater genetic distinctiveness of C. angustifolia, even when growing in sympatry with the other two species, is in accordance with ancestral lineage divergence during the late Miocene (following Koenen et al., 2015). As a consequence, interspecific reproductive isolation seems to be incomplete between C. balansae and C. saltensis as shown by the common widespread H2 cpDNA haplotype shared by the two species. In a recent study, where C. balansae and C. saltensis occur in sympatry, genetically pure samples of either species were found together with hybrids and introgressant forms in several populations (Zelener et al., 2016). Nonetheless, shared cpDNA sequences at the northern-most studied location where the three species grow in sympatry highlights the potential for hybridization among the three species. While it is unknown if pollen from one species of Cedrela can pollinate other species, the lack of chloroplast DNA differentiation and thus chloroplast sharing may be due to deficient reproductive barriers by relatively recent speciation or secondary contact that occurred under particular ecological and climatic conditions along altitudinal gradients. Previous phylogenetic studies of the Meliaceae based on cpDNA resulted in polytomies (Muellner et al., 2009) suggesting that hybridization may be common.
Chloroplast sharing, i.e. the lack of interspecific polymorphism at cpDNA, may be due to convergent evolution, incomplete lineage sorting, and/or reticulate evolution (Acosta & Premoli, 2010). Sequence convergence would be highly unlikely. The high potential for interspecific gene flow in plants (Rieseberg & Soltis, 1991) may result in chloroplast capture. This is where the cytoplasm of one species is replaced by that of another species through hybridization/introgression (Fehrer et al., 2007). Chloroplast capture occurs frequently in species with sympatric distributions and reproductive compatibility (Acosta & Premoli, 2010). Thus, in areas with range overlap and under similar ecological settings, Cedrela species may produce hybrids whereas under contrasting environmental conditions, e.g. elevation, they may diverge by ecological selection even under potential interspecific gene flow. Hence, varying conditions along elevation gradients have probably driven ecological divergence in Cedrela.
Appendix. Allele frequencies of three Cedrela species at eight isozyme loci. CA: C. angustifolia, CB: C. balansae, CS: C. saltensis.
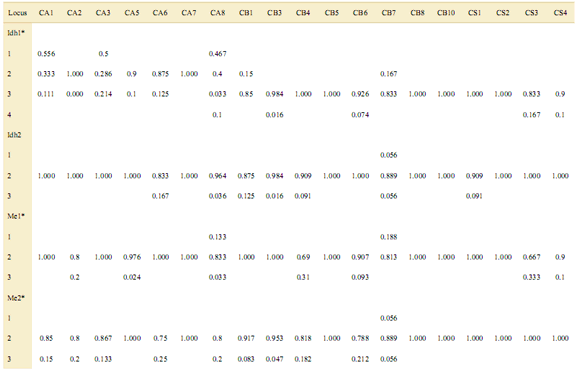
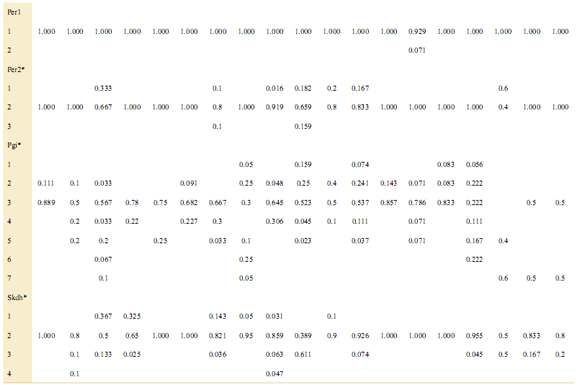
*p<0.05 chi-square test of heterogeneity of allele frequencies along populations.
ACKNOWLEDGMENTS
Ezequiel Balducci, Eugenia Enrrico, Roberto Cáceres for sample collection. Martina Fernández for collaboration in the laboratory. Romina Vidal-Russel for assistance in statistical analyses. Silvia Pacheco and Karina Buzza for the design of figures 1 and 4. This research was carried out in the framework of the Re-ForLan project funded by the European Commission (FP6-2004-INCODEV-3 PROP no. 032132).
BIBLIOGRAPHY
1. Acosta, M. C. & A. C. Premoli. 2010. Evidence of chloroplast capture in South American Nothofagus (subgenus Nothofagus, Nothofagaceae). Molecular Phylogenetics and Evolution 54: 235-242. DOI: 10.1016/j.ympev.2009.08.008 [ Links ]
2. Akaike, H. 1973. Information theory and an extension of the maximum likelihood principle, in Petrov B. N. & F. Caski (eds.), [Proceedings of the 2nd International Symposium on Information Theory]. Budapest: Akadémiai Kiado, 267-281. [ Links ]
3. Cavers, S.; A. Telford, F. Arenal Cruz, A. J. Pérez Castañeda, R. Valencia, C. Navarro, A. Buonamici & A. J. Lowe. 2013. Cryptic species and phylogeographical structure in the tree Cedrela odorata L. throughout the Neotropics. Journal of Biogeography 40: 732-746. DOI: 10.1111/jbi.12086 [ Links ]
4. Coyne, J. A. & H. A. Orr. 2004. Speciation. Suderland, MA: Sinauer. [ Links ]
5. Demesure B.; N. Sodzi & R. J. Petit. 1995. A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Molecular Ecology 4: 129-131. DOI: 10.1111/j.1365-294X.1995.tb00201.x [ Links ]
6. Dumolin, S.; B. Demesure & R. J. Petit. 1995. Inheritance of chloroplast and mitochondrial genomes in pedunculate oak investigated with an efficient PCR method. Theoretical and Applied Genetics 91: 1253-1256. DOI: 10.1007/BF00220937 [ Links ]
7. Fehrer, J.; B. Gemeinholzer, J. Chrtek & S. Bräutigam. 2007. Incongruent plastid and nuclear DNA phylogenies reveal ancient intergeneric hybridization in Pilosella hawkweeds (Hieracium, Cichorieae, Asteraceae). Molecular Phylogenetics and Evolution 42: 347-361. DOI: 10.1016/j.ympev.2006.07.004 [ Links ]
8. Garcia-Gonçalvez, M.; R. S. Silva, M. A. Carniello, J. W. Veldman, A. A. B. Rossi & L. O. de Oliveira. 2011. Molecular evidence of cryptic speciation, historical range expansion, and recent intraspecific hybridization in the Neotropical seasonal forest tree Cedrela fssilis (Meliaceae). Molecular Phylogenetics and Evolution 61(3): 639-649. DOI: 10.1016/j.ympev.2011.08.026 [ Links ]
9. Goudet, G. 1995. FSTAT (Version 1.2): A Computer Program to Calculate F-Statistics. Journal of Heredity 86(6): 485-486. [ Links ]
10. Grau, A.; M. A. Zapater & R. A. Neumann. 2006. Botánica y distribución del género Cedrela en el noroeste de Argentina, en Pacheco S. & A. Brown (eds.), Ecología y producción de cedro (género Cedrela) en las Yungas australes: 19-30. Tucumán: Ediciones del Subtrópico. [ Links ]
11. Hamilton, M. B. 1999. Four primer pairs for the amplification of chloroplast intergenic regions with intraspecific variation. Molecular Ecology 8(3): 521-523. [ Links ]
12. Koch, M. A.; C. Dobeš & T. Mitchell-Olds. 2003. Multiple hybrid formation in natural populations: concerted evolution of the internal transcribed spacer of nuclear ribosomal DNA (ITS) in North American Arabis divaricarpa (Brassicaceae). Molecular Biology and Evolution 20(3): 338-350. DOI: 10.1093/molbev/msg046
13. Köcke, A. V. 2015. Spatio-temporal evolution of Cedrela (Meliaceae): climatic niche dynamics, phylogeography and taxonomy. PhD Thesis, Johann Wolfgang Goethe Univesity, Germany. [ Links ]
14. Koecke, A.; A. Muellner-Riehl, T. D. Pennington, G. Schorr & J. Schnitzler. 2013. Niche evolution through time and across continents: the story of the neotropical Cedrela (Meliaceae). American Journal of Botany 100: 1800-1810. DOI: 10.3732/ajb.1300059 [ Links ]
15. Koenen, E. J. M.; J. J. Clarkson, T. D. Pennington & L. W. Chatrou. 2015. Recently evolved diversity and convergent radiations of rainforest mahoganies (Meliaceae) shed new light on the origins of rainforest hyperdiversity. New Phytologist 207(2): 327-339. DOI: 10.1111/nph.13490 [ Links ]
16. Librado, P. & J. Rozas. 2009. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25(11): 1451-1452. DOI: 10.1093/bioinformatics/btp187 [ Links ]
17. Malizia L. R.; C. Blundo, S. E. Pacheco. 2006. Diversidad, estructura y distribución de bosques con cedro (Cedrelas pp., Meliaceae) en noroeste de Argentina y sur de Bolivia, En: Pacheco, S. & A. Brown (eds.), Ecología y producción de cedro (género Cedrela) en las Yungas australes: 83-104. Tucumán: Ediciones del Subtrópico, [ Links ]
18. Mayr, E. 1947. Ecological factors in speciation. Evolution 1(4): 263–288. DOI: 10.2307/2405327
19. Mitton. J. B.; Y. B. Linhart, K. B. Sturgeon & J. L. Hamrick. 1979. Allozyme polymorphisms detected in mature needle of ponderosa pine. Journal of Heredity 70: 86-89. [ Links ]
20. Muellner, A. N.; R. Samuel, M. W. Chase, C. M. Pannell & H. Greger. 2005. Aglaia (Meliaceae): an evaluation of taxonomic concepts based on DNA data and secondary metabolites. American Journal of Botany 92(3): 534-543. DOI: 10.3732/ajb.92.3.534 [ Links ]
21. Muellner, A. N.; T. D. Pennington & M. W. Chase. 2009. Molecular phylogenetics of neotropical Cedreleae (mahogany family, Meliaceae) based on nuclear and plastid DNA sequences reveal multiple origins of “Cedrela odorata”. Molecular Phylogenetics and Evolution 52(2): 461-469. DOI: 10.1016/j.ympev.2009.03.025
22. Muellner, A. N.; T. D. Pennington, A. V. Koecke & S. S. Renner. 2010. Biogeography of Cedrela (Meliaceae, Sapindales) in Central and South America. American Journal of Botany 97(3): 511-518. DOI: 10.3732/ajb.0900229 [ Links ]
23. Muir, G.; C. C. Fleming & C. Schlötterer. 2001. Three divergent rDNA clusters predate the species divergence in Quercus petraea (Matt.) Liebl. and Quercus robur L. Molecular Biology and Evolution 18(2): 112-119. [ Links ]
24. Navarro, C.; S. Ward & M. Hernández. 2002. The tree Cedrela odorata (Meliaceae): a morphologically subdivided species in Costa Rica. International Journal of Tropical Biology and Conservation 50: 21-29. [ Links ]
25. Nylander, J. A. 2004. MrModelTest. Program distributed by the author. Evolutionary Biology Centre, Uppsala University, Uppsala. Available on: http://www.abc.se/~nylander/ [ Links ]
26. Orr, M. R. & T. B. Smith. 1998. Ecology and speciation. Trends in Ecology and Evolution 13(2): 502-506. [ Links ]
27. Pennington, T. P. & A. N. Muellner. 2010. A monograph of Cedrela (Meliaceae). England: DH Books. [ Links ]
28. Posada, D. & K. A. Crandall. 1998. Modeltest: testing the model of DNA substitution. Bioinformatics Applications Note 14(9): 817-818. DOI: 10.1093/bioinformatics/14.9.817 [ Links ]
29. Poulik, M. D. 1957. Starch gel electrophoresis in a discontinuous system of buffers. Nature (London) 180: 1477-1479. DOI: 10.1038/1801477a0 [ Links ]
30. Premoli, A. C., C. P. Souto, S. Trujillo A., R. del Castillo, P. Quiroga, T. Kitzberger, Z. Gomez O., M. Arbetman, L. R. Malizia, A. Grau, R. Rivera G. & A.C. Newton. 2011. Impact of forest fragmentation and degradation on patterns of genetic variation and its implication for forest restoration. In: Principles and Practice of Forest Landscape Restoration. Case studies from the drylands of Latin America (eds. A.C. Newton & N. Tejedor G.) International Union for the Conservation of Nature. [ Links ]
31. Ranker, T. A.; C. H. Haufler, P. S. Soltis & D. E. Soltis. 1989. Genetic evidence for allopolyploidy in the neotropical fern Hemionitis pinnatifida (Adiantaceae) and the reconstruction of an ancestral genome. Systematic Botany 14(4): 439-447. DOI: 10.2307/2418989 [ Links ]
32. Rieseberg, L. H. & D. E. Soltis. 1991. Phylogenetic consequences of cytoplasmic gene flow in plants. Evolutionary Trends in Plants 5(1): 65-84. [ Links ]
33. Ronquist, F. & J. P. Huelsenbeck. 2003. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12): 1572-1574. DOI: 10.1093/bioinformatics/btg180 [ Links ]
34. Rundle, H. D. & P. Nosil. 2005. Ecological speciation. Ecology Letters 8: 336-352. [ Links ]
35. Schluter, D. 2001. Ecology and the origin of species. Trends in Ecology & Evolution, 16(7): 372-380. [ Links ]
36. Taberlet, P.; L. Gielly, G. Pautou & J. Bouvet. 1991. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Molecular Biology 17(5): 1105-1109. [ Links ]
37. Tamura, K.; J. Dudley, M. Nei & S. Kumar 2007. MEGA4 : Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Molecular Biology and Evolution 24: 1596-1599. DOI: 10.1093/molbev/msm092 [ Links ]
38. Wright, S. 1965. The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19(3): 358-420. DOI: 10.2307/2406450 [ Links ]
39. Wright, S. 1978. Evolution and the genetics of populations, vol. 4: Variability within and among natural populations. Chicago: University of Chicago Press. [ Links ]
40. Yeh, F.C. & T. J. B. Boyle. 1997. Population genetic analysis of co-dominant and dominant markers and quantitative traits. Belgian Journal of Botany 129: 157-163. [ Links ]
41. Zapater, M. A.; E. M. del Castillo, & T. D. Pennington. 2004. El género Cedrela (Meliaceae) en la Argentina. Darwiniana 42: 347-356. [ Links ]
42. Zelener, N.; D. Tosto, L. O. de Oliveira, M. A. Soldati, M. V. Inza, & L. F. Fornes. 2016. Molecular evidence of hybrid zones of Cedrela (Meliaceae) in the Yungas of Northwestern Argentina. Molecular Phylogenetic and Evolution 102: 45-55. [ Links ]














