Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO  uBio
uBio
Compartir
Revista argentina de microbiología
versión impresa ISSN 0325-7541versión On-line ISSN 1851-7617
Rev. argent. microbiol. v.41 n.2 Ciudad Autónoma de Buenos Aires abr./jun. 2009
Identification of the source of histoplasmosis infection in two captive maras (Dolichotis patagonum) from the same colony by using molecular and immunologic assays
M. R. Reyes-Montes1, G. Rodríguez-Arellanes1, A. Pérez-Torres2, A. G. Rosas-Rosas3, A. Parás-García3, C. Juan-Sallés4, M. L. Taylor1 *
Departamentos de 1Microbiología-Parasitología y
2Biología Celular, Facultad de Medicina, Universidad Nacional Autónoma de México, México DF, (04510) Mexico;
3Departamento de Veterinaria, Africam Safari, 11 Oriente 2407 (Col. Azcárate), Puebla, Puebla, (72007) México;
4ConZOOlting Wildlife Management, La Canvella 1, Nau 1, (08445) Samalús, Spain
*Correspondence. E-mail: emello@servidor.unam.mx
ABSTRACT
Histoplasma capsulatum was isolated from the spleen of a first infected mara (Dolichotis patagonum) and from a second mara's liver and adrenal gland, both in the same colony at the Africam Safari, Puebla, Mexico. Studies of H. capsulatum isolates, using nested-PCR of a 100-kDa protein coding gene (Hcp100) fragment and a two-primer RAPDPCR method, suggest that these isolates were spreading in the environment of the maras' enclosure. By using a Dot- ELISA method, sera from mice inoculated with three homogenates of soil samples from the maras' enclosed space developed positive brown spot reactions to a purified H. capsulatum antigen, which identified the probable source of the maras' infection.
Key words: Histoplasma capsulatum; Maras; PCR; RAPD-PCR; Dot-ELISA; Infection source
RESUMEN
Identificación de la fuente de infección de histoplasmosis de dos maras (Dolichotis patagonum) cautivas procedentes de la misma colonia, utilizando ensayos moleculares e inmunológicos. Histoplasma capsulatum fue aislado del bazo de una primera mara (Dolichotis patagonum) infectada y del hígado y la glándula suprarrenal de un segundo ejemplar, ambos de la misma colonia en el Africam Safari, Puebla, México. Los estudios de los aislamientos de H. capsulatum mediante PCR anidada para un fragmento del gen Hcp100 que codifica una proteína de 100 kDa y RAPD-PCR empleando doble iniciador sugieren que estos aislamientos estaban dispersos en el ambiente del refugio de las maras. Los sueros de ratones inoculados con tres homogenatos de muestras de suelo del refugio desarrollaron reacciones positivas a un antígeno purificado de H. capsulatum (manchas color castaño oscuro) por el método de Dot-ELISA; con lo cual se identificó la probable fuente de infección de las maras.
Palabras clave: Histoplasma capsulatum; Maras; PCR; RAPD-PCR; Dot-ELISA; Fuente de infección
The pathogenic fungus Histoplasma capsulatum var. capsulatum, the etiologic agent of the systemic mycosis histoplasmosis, usually occurs in geographic areas where tropical climates prevail. It is present either in confined spaces where bat guano is abundant or in open spaces where bird droppings are frequently found. Infection is acquired from the environment by inhalation of infective mycelial-phase propagules that are aerosolized (11).
Bats are probably the major fungal reservoir that contributes to maintain this parasite in natural foci. Histoplasmosis has been described in domestic and freeranging wildlife animals. Occasionally, it has been documented in captive animals, including a sea otter (Enhydra lutris) (3) and an owl monkey (Aotus nancymai) (12). Disseminated histoplasmosis in captive maras (Dolichotis patagonum), also called Patagonian hares, at the Africam Safari, in the state of Puebla, Mexico, has been described elsewhere (8, 9). We herein report molecular and immunologic findings that support the identification of the source of infection in the maras' enclosure, which were probably associated with two histoplasmosis cases in captive maras from the same colony at the Africam Safari, Mexico.
Three isolates from maras were used: the EH-558B from the spleen belonging to case 1 and EH-574A and EH-574H from the adrenal gland and liver of case 2, respectively. Isolates were identified by morphological and antigenic procedures as described by Taylor et al. (10). One isolate (EH-406) from a naturally infected bat and another one from an Argentinean patient (01558) were processed as controls in different molecular assays. The mycelial phase of each H. capsulatum isolate was maintained at 26 ºC in brain-heart-infusion agar (BHI-agar) (Bioxón, México, DF).
DNA samples were extracted from the three maras' H. capsulatum isolates and from the two control isolates. Whole-cell DNA isolation was performed as described elsewhere by Reyes-Montes et al. (6).
Nested-PCR was performed as described by Bialek et al. (1) with minor modifications by Taylor et al. (10). DNA samples from the studied H. capsulatum isolates were processed together with a DNA sample of Sporothrix schenckii, used as negative control of heterologous fungal strain. Two sets of primers (Operon Technologies Inc., Alameda, CA) were used, corresponding to a fragment of the gene encoding for a 100-kDa protein (Hcp100) unique to H. capsulatum (10). The outer primer set, Hc I and Hc II, delimits a 391-nucleotide sequence of the gene (1). The inner primers, Hc III and Hc IV, delimit a specific 210- nucleotide sequence (1). DNA amplification and the electrophoresis of the PCR products were performed according to Taylor et al. (10).
The two-primer RAPD-PCR assay was performed as described by Hu et al. (2) with modifications by Taylor et al. (10). The amplification patterns of different isolates were analyzed to obtain an estimation of similarity for each pair of isolates. The similarity was calculated by using the Jaccard coefficient (5). A dendrogram was constructed based on the similarity matrix and through the unweighted pair-group method with arithmetic averages (UPGMA). Multivariate statistical methods were performed with the NTSYS-PC program (version 2.0; Exeter Software) (7).
To detect the maras' sources of infection, sera from mice inoculated with homogenates from some environmental samples in different places of the maras' enclosure at the Africam Safari (Table 1) were processed through Dot-ELISA (4, 10). Criteria to define positive reaction in sera from tested mice were standardized by taking into account that two out of three mice sera revealed brown spots in the assay.
Table 1. Dot-ELISA from sera of mice inoculated with different environmental samples collected from the maras' enclosure at the Africam Safari, Puebla, Mexico

(1)Sera from mice inoculated with homogenates of the environmental samples were harvested and tested at 10 and 17 days after mice inoculation. Dot-ELISA was performed by using a purified DPPC-Histo molecule as specific H. capsulatum antigen (10). Samples S-190, S-191, and S-197 were collected in distinct places of the maras' open shelter. Samples S-195 and S-196 were collected from different burrows in the middle of the maras' enclosure.
H. capsulatum isolates from the maras' samples were previously identified by morphological and antigenic findings and are now preserved in the H. capsulatum Culture Collection of our laboratory.
Nested-PCR products targeting the Hcp100 protein gene showed that all H. capsulatum isolates from the infected maras shared the same bands (391 and 210 bp in the first and nested amplifications, respectively) with two H. capsulatum isolates tested as controls. DNA of S. schenckii was not amplified (Figure 1 a and b).

Figure 1. Nested-PCR of H. capsulatum DNA samples from infected maras. The assay was performed with two sets of fungal-specific primers of the 100-kDa protein gene of H. capsulatum. PCR products were resolved by electrophoresis in 1.5% agarose followed by ethidium bromide-staining. First (a) and nested (b) PCR reactions. Lanes refer to each processed H. capsulatum isolate. The S. schenckii (Ss) DNA was used as negative control of heterologous strain, and C (-) as negative control of the system. M- 123 bp DNA ladder molecular size marker. Abbreviations: AR- Argentina; MX/PL- Mexico/Puebla; GT- Guatemala.
The RAPD-PCR analysis distinguished the three maras' H. capsulatum isolates from the two different H. capsulatum isolates used as controls. The dendrogram in Figure 2 formed a highly related group of maras' H. capsulatum isolates, including the isolates from case 2 (EH-574A and 574H) with a 1.0 similarity and the isolate from case 1 (EH-558B) that showed a 0.84 similarity with those from case 2. In contrast, a less related group with a 0.36 similarity was formed by one isolate from a naturally infected bat captured in Puebla (EH-406), and another one from an Argentinean human clinical case (01558).
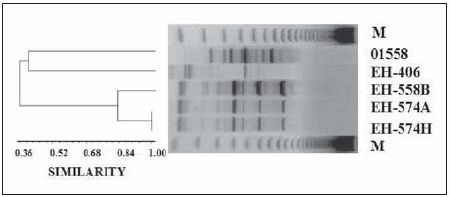
Figure 2. Polymorphic DNA profiles of the maras' H. capsulatum isolates. Three isolates from both maras (EH-558B, EH-574A, and EH-574H), one isolate from a naturally infected bat (EH- 406), and another isolate from an Argentinean patient (01558) were processed, using 10 ng of DNA template in a two-primer RAPD-PCR assay. The PCR products were resolved by electrophoresis in 1.5% agarose followed by ethidium bromide-staining. The dendrogram was constructed based on the UPGMA algorithm and the similarity between each pair of isolates was calculated by the Jaccard coefficient. M- 123 bp DNA ladder molecular size marker.
The presence of anti-H. capsulatum antibodies in sera of most mice at 10 and 17 days after inoculation with samples S-190, S-191, and S-197 suggests that the source of the maras' infection was associated with bat and bird guano collected from places related to the maras' enclosure, where both types of guano had accumulated near the maras' food container (Table 1).
Although H. capsulatum isolates from the mara samples were undoubtedly identified by morphological and antigenic findings, the molecular studies of these isolates were an important support to relate H. capsulatum isolates sampled from other sources and geographic origins. At the same time, the infection source at the maras' enclosure was immunologically monitored by detecting positive reaction to H. capsulatum antigen in sera of mice inoculated with different environmental samples, including from their burrows.
The nested-PCR assay established that the mara isolates shared the same bands in all H. capsulatum isolates studied, which supports their molecular recognition. Besides, the RAPD-PCR findings revealed, through the dendrogram based on the UPGMA analysis, that the mara isolates have high similarity in their polymorphic DNA bands, which differed from the DNA patterns of the two H. capsulatum control isolates processed. Thus, RAPDPCR results suggest that the mara isolates shared the same source of infection.
The presence of H. capsulatum in the maras' enclosure was well-demonstrated by antibody production in mice inoculated with the environmental samples.
This work details infection data from two captive maras with disseminated histoplasmosis, regarding molecular findings of their H. capsulatum isolates and the immunological identification of their respective infection sources.
Acknowledgements: The authors thank Ingrid Mascher for editorial assistance.
1. Bialek R, Feucht A, Aepinus C, Just-Nubling G, Robertson VJ, Knobloch J, et al. Evaluation of two nested PCR assays for detection of Histoplasma capsulatum DNA in human tissue. J Clin Microbiol 2002; 40: 1644-7. [ Links ]
2. Hu J, van Eysden J, Quiros CF. Generation of DNA-based markers in specific genome regions by two-primer RAPD reactions. In: PCR Methods and Applications. Cold Spring Harbor, Cold Spring Harbor Laboratory Press, 1995, p. 346-51. [ Links ]
3. Morita T, Kishimoto M, Shimada A, Matsumoto Y, Shindo J. Disseminated histoplasmosis in a sea otter (Enhydra lutris). J Comp Pathol 2001; 125: 219-23. [ Links ]
4. Papas MG, Hajkowski R, Hockmeyer WT. Dot-enzyme linked immunosorbent assay (Dot-ELISA): A micro technique for the rapid diagnosis of visceral leishmaniasis. J Immunol Methods 1983; 64: 205-14. [ Links ]
5. Real R, Vargas JM. The probabilistic basis of Jaccard's index of similarity. Syst Biol 1996: 45: 380-5. [ Links ]
6. Reyes-Montes MR, Bobadilla-Del Valle M, Martínez-Rivera MA, Rodríguez-Arellanes G, Maravilla E, Sifuentes- Osornio J, et al. Relatedness analyses of Histoplasma capsulatum isolates from Mexican patients with AIDS-associated histoplasmosis by using histoplasmin electrophoretic profiles and randomly amplified polymorphic DNA patterns. J Clin Microbiol 1999; 37: 1404-8. [ Links ]
7. Rohlf FJ. NTSYS-pc. Numerical taxonomy and multivariate analysis system, version 2.0. Applied Biostatistics, New York, 1998. [ Links ]
8. Rosas-Rosas AG, Juan-Sallés C, Garner MM. Pathological findings in a captive colony of maras (Dolichotis patagonum). Vet Rec 2006; 158: 727-31. [ Links ]
9. Rosas-Rosas A, Juan-Sallés C, Rodríguez-Arellanes G, Taylor ML, Garner MM. Disseminated Histoplasma capsulatum var. capsulatum infection in a captive mara (Dolichotis patagonum). Vet Rec 2004; 155: 426-8. [ Links ]
10. Taylor ML, Ruíz-Palacios GM, Reyes-Montes MR, Rodríguez-Arellanes G, Carreto-Binaghi LE, Duarte-Escalante E, et al. Identification of the infection source of an unusual outbreak of histoplasmosis, in a hotel in Acapulco, state of Guerrero, Mexico. FEMS Immunol Med Microbiol 2005; 45: 435-41. [ Links ]
11. Tewari R, Wheat LJ, Ajello L. Agents of histoplasmosis. In: Ajello L, Hay RJ, editors. Medical Mycology. Topley& Wilson's, Microbiology and Microbial Infections. 9th edition. New York, Arnold and Oxford University Press, 1998, p. 373-407. [ Links ]
12. Weller RE, Dagle GE, Malaga CA, Baer JF. Hypercalcemia and disseminated histoplasmosis in an owl monkey. J Med Primatol 1990; 19: 675-80. [ Links ]
Recibido: 12/08/08
Aceptado: 11/11/08














