Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO  uBio
uBio
Compartir
Revista argentina de microbiología
versión impresa ISSN 0325-7541
Rev. argent. microbiol. vol.42 no.4 Ciudad Autónoma de Buenos Aires oct./dic. 2010
CLINICAL MICROBIOLOGY AND INFECTIOUS DISEASES
Evaluation of an immunomagnetic capture method followed by PCR to detect Mycobacterium bovis in tissue samples from cattle
S. G. Garbaccio1, A. A. Cataldi2*
1Instituto de Patobiología.
2Instituto de Biotecnología. CICVyA. Instituto Nacional de Tecnología Agropecuaria (1686), Hurlingham, Argentina
*Correspondence. E-mail: acataldi@cnia.inta.gov.ar
ABSTRACT
Tuberculosis is one of the most important infectious diseases worldwide. Mycobacterium bovis is the causative agent of bovine tuberculosis (BTB), an important animal pathogen with public health implications as it is a zoonosis. Currently, the diagnosis of BTB is based on the caudal fold test of the Tuberculin Skin Test (TST). Post-mortem bacterial culture is carried out to confirm the diagnosis, and then specific biochemical tests are performed for the characterization of the etiologic agent. Culture takes at least 4 to 8 weeks to develop. The diagnosis by molecular tests such as PCR can provide fast and reliable results, significantly decreasing the time of confirmation (from two months to two days), thus allowing the possibility of taking control actions to prevent the spread of the disease in herds. In this work the use of an immunomagnetic separation capture followed by PCR (IMS-PCR) based on the IS6110 element showed a detection threshold corresponding to 10 CFU in M. bovis-spiked PBS. In the case of infected bovine fresh tissues, after five replicates, the minimum value of detection was 1000 CFU in 100% of the trials (5/5). This paper attempts to provide a sensitive, rapid and specific technique for the diagnosis of bovine tuberculosis, and opens up the possibility of a direct application in the control and eradication of this cattle disease.
Key words: Mycobacterium bovis; Immunomagnetic capture; PCR; Cattle
RESUMEN
Evaluación de un método de captura inmunomagnética seguida de PCR para la detección de Mycobacterium bovis en tejidos de ganado. La tuberculosis es una de las enfermedades infecciosas más importantes. Mycobacterium bovis es el agente causal de la tuberculosis bovina (TBB), un patógeno animal y zoonótico. En la actualidad, el diagnóstico de TBB se basa en la prueba intradérmica de la tuberculina. El cultivo bacteriano post mortem se lleva a cabo para confirmar el diagnóstico y a continuación se realizan pruebas bioquímicas específicas para la caracterización del agente etiológico. El cultivo bacteriano toma por lo menos 4 a 8 semanas para su desarrollo. El diagnóstico mediante pruebas moleculares como PCR puede proporcionar resultados rápidos y robustos, con un considerable acortamiento hasta la confirmación del diagnóstico (de 2 meses a 2 días). En este trabajo, el uso de captura inmunomagnética seguida de PCR (IMS-PCR) dirigida al elemento IS6110 mostró un umbral de detección correspondiente a 10 UFC en M. bovis diluido en PBS. En el caso de tejidos bovinos inoculados experimentalmente después de 5 réplicas, el valor mínimo de detección fue de 1000 UFC en el 100% de los ensayos. Este artículo aspira a proporcionar una técnica sensible, rápida y específica para el diagnóstico de la tuberculosis bovina, con el fin de abrir la posibilidad de una aplicación directa en el control y la erradicación de esta enfermedad en el ganado.
Palabras clave: Mycobacterium bovis; Captura inmunomagnética; PCR; Ganado
Introduction
Bovine tuberculosis is a chronic infectious disease and also a zoonosis which affects cattle and other species of domestic or wild animals. Mycobacterium bovis, the causative agent of bovine tuberculosis, is a slow growing mycobacterium that belongs to the Mycobacterium tuberculosis complex group. This is a closely related group that is pathogenic both for humans and animals. This taxonomic complex comprises the following species: M. tuberculosis, M. bovis, which includes the BCG strain (Bacillus Calmette Guerin, attenuated mutant of M. bovis), Mycobacterium africanum (heterogeneous group responsible for human tuberculosis in equatorial Africa), Mycobacterium microtii and Mycobacterium canetti. Recently, M. tuberculosis subsp. caprae (2) and Mycobacterium pinnipedii subsp. nov. (3) have been incorporated into this group. The M. bovis genome which has been recently sequenced is 99.5% identical to that of M. tuberculosis H37Rv, but smaller due to deletions (5).
The progress of control and eradication programs in many countries, coupled with the widespread implementation of pasteurization, has led to the reduction in the incidence of human infection by M. bovis. In humans, the infection occurs primarily through inhalation of aerosols from coughing cattle in abattoirs, or by the consumption of contaminated non-pasteurized milk (7, 17). However, in many countries where the disease in cattle is endemic, M. bovis still persists in humans.
In a work carried out in Argentina, 2.2% of the patients with tuberculosis in the province of Santa Fe and 1% of the patients in the province of Buenos Aires (4) presented M. bovis as its causative agent. In most cases, there was an association with occupational exposure.
Although bacteriological culture is considered the gold standard test for diagnostic confirmation, it takes two months to confirm diagnosis. The direct smear from sputum examination is considered a fast and easy test for the diagnosis of human tuberculosis. However, it is not useful in bovine TB because it lacks sensitivity for BTB diagnosis (22).
Several studies have demonstrated the potential of PCR in clinical samples for the detection of BTB, with results comparable to those obtained from bacterial culture, although in shorter times: 2-3 days for PCR versus 4-8 weeks for bacterial cultures (1, 11, 26). Different methodologies have been described for extracting and recovering DNA prior to PCR, ranging from boiling and centrifugation, boiling and cold, ultrasound, bead beating, enzymatic digestion, immunomagnetic beads, phenol-chloroform extraction, commercial kits for lysis and purification, and the combination of several of them (1, 24, 26).
The present work aims at developing a PCR technique in fresh tissues, using immunomagnetic separation (IMS) as a method to capture bacteria. Although this IMS-PCR combination has been tested in mycobacteria present in different bovine clinical materials such as feces or milk (1, 9), there are no reports of its use in tissues. Therefore, this technique could help in the diagnosis of special cases, for example, in the event of an outbreak where immediate intervention is needed or for the rapid definition of a case having lesions compatible with bovine tuberculosis.
MATERIALS AND METHODS
Seed lot culture
A seed culture was obtained in liquid medium (Middlebrook 7H9 OADC) for growth of M. bovis AN5 (reference strain), incubated at 37 °C for four weeks. Prior to counting, and taking into account that a characteristic of mycobacteria is "clumping", different methodologies were established to evaluate which could best separate these formations before loading them on Petri dishes and counting. Three methods used were: Passage through a 5μm Millipore filter using a tuberculin syringe type (a); grinding in a potter homogenizer with 1 ml of culture placed in the container, making 25 up and down movements with the pestle (b) and repeated tuberculin needle passage: 1 ml of culture placed in a 15 ml tube, and 30 up and down movements using a tuberculin syringe and needle to disintegrate the mycobacterial accumulation (c).
To count CFU, 1ml aliquots were taken from the seed lot and disaggregated as described. Then, serial 10 fold dilutions were poured in duplicate on Petri plates with Middlebrook 7H10 solid medium and incubated at 37 °C until colony development and CFU determination at 35 days of culture.
Dilutions of M. bovis seed lot culture
Serial 10-fold dilutions of the seed lot culture were prepared by vigorous mixing as described in method (c) in the previous section, using sterile phosphate buffer solution (PBS) with 0.05% (v/v) Tween 20. Dilutions were aliquoted in 1.5ml tubes and kept at -20 °C until inoculation.
Inoculation of tissues
Various organs (retropharyngeal, mediastinal, tracheobronchial, mesenteric and hepatic lymph nodes, lungs, and liver), from animals free of bovine tuberculosis were used. A pool was made of each organ and placed in a plastic bag with 100 ml of sterile 1XPBS. The mixture was homogenized using a mechanical grinder (Basic Masticator type 470, IUL Instruments), for 6 beats (1.5 minutes each) until the formation of a homogenate. This material was aliquoted and kept at -20 °C until experimental inoculation.
Polyclonal antibody (PA)
PA was provided by the reference laboratory of the University of Colorado-USA (http://www.cvmbs.colostate.edu/microbiology/tb/material.html), and produced in rabbit inoculated with M. tuberculosis H37Rv (complete cell lysate), and the original titer in Western blot was reported to be 1:24,000. The lyophilized PA was reconstituted and the titer was tested in Western blot against a M. bovis AN5 cell extract (E) and culture supernatant (SB). Both materials were subjected to PAGE-SDS and four decreasing dilutions of polyclonal sera were used: 1:100, 1,000, 10,000 and 100,000. The protein concentration of sera was quantified against a standardized curve of albumin by the Bradford method, resulting in a total concentration of 107.4 mg/ml, and then diluted (1:3) and aliquoted.
Evaluation of the optimal concentration of polyclonal sera to use in the preparation of conjugated beads
To assess the efficiency of the beads-PA conjugate we worked with 100 and 1,000 CFU, varying the antibody concentration. The beads were used at a fixed number of 12 μl per milliliter of sample, as described by Antognoli et al. (1), who used a concentration of 8 μg of PA in 120 μl of beads, (i.e. 0.066 mg of PA per ml of beads). The following scheme was applied in duplicate: dilution 1 (0.07 μg of PA/μl beads), dilution 2 (0.7 µg/µl), dilution 3 (7 µg/µl) and dilution 4 (15 µg/µl). We used fixed values of mycobacteria: 100 and 1,000 CFU diluted in 1 ml of PBS 1X containing 0.05% of Tween 20. The optimal concentration of PA was determined by the minimal detection capacity of IMS-PCR.
Determination of the optimal binding time of Mycobacterium to the conjugate (beads+polyclonal)
As described by Antognoli et al (1), the infected material was incubated with the conjugate for 1.5 h. Taking this value as a reference, incubations were carried out in duplicate at 0.5 h, 1, 1.5 and 2 h. The two CFU (100 and 1,000) diluted in 1ml of 1X PBS containing 0.05% Tween 20 were used. This step was first evaluated by specific Ziehl Neelsen staining and then by IMS-PCR.
IMS Protocol
Paramagnetic beads (Spherotech, Dynal) with attached goat anti-rabbit IgG were incubated with anti-M. tuberculosis H37Rv lysate rabbit serum. The protocol previously described (1), was adapted with minor modifications. The PA was conjugated to the beads for 1 h by incubation in constant motion at room temperature. Then, 12µl of conjugate was added to 1ml of sample and incubated for 1.5 h in constant motion at room temperature. The tubes containing samples and beads were left in a magnetic field (Magnetic particle concentrator and simple mixer-Dynal MPC-S) for 30 minutes. The remaining liquid was removed carefully with a tip, leaving 50µl of liquid in the tube. Then, 1ml of 0.05%, PBS Tween was added, and the tube was placed back into the magnet for 10 minutes. This operation was repeated three times, until the liquid became transparent. Finally, the beads were resuspended in 100 µl of sterile distilled water and subjected to lysis for 15 min at 100 °C for the release of mycobacterial DNA. After a centrifugation (13,000 rpm for 5 min) to pellet the beads, the supernatant was used as template for PCR.
The amplification conditions were performed according to Zumarraga et al. (25). A region of the IS6110 insertion sequence was amplified using primers previously described (8-20), generating a 245-bp amplification product. Master Mix consisted of the following reagents: Taq buffer 5 μl, MgCl2 2.5 mm, dNTP 0.2 mM, INS1 (cgtgagggcatcgaggtggc) g 1 μM and INS2 (cgtaggcgtcggtgacaaa) 1 μM, 34.8 µl sterile water, Taq polymerase (1.25 U) 0.2 μl, and DNA template 2 μl. The final volume was 50 μl. For the amplification process a thermocycler MJ PTC100 was used with the following program: 96 °C for 3 min, 10 cycles of 96°C for 1 min, 72 °C for 1 min decreasing 1 °C / cycle (Touch Down), 72 °C for 1 min, and 30 cycles of 96 °C for 1 min, 65 °C for 1 min, 72 °C for 2 min, 72 °C for 8 min. The product was detected by electrophoresis in 2% agarose gels at a constant voltage (120 V).
Detection limit of IMS-PCR in PBS
Duplicate decreasing dilutions of M. bovis AN5 DNA were performed, from an initial concentration of 3 x 105 followed by five successive ten fold dilutions in PBS tween 0.5% to a final volume of 100 µl. The tubes were placed in a water bath at 100 °C for 20 min and the material used as a template for PCR. This operation was repeated five times on different days.
Detection limit using IMS-PCR in bovine tissues
Tubes containing 1 ml of tissue homogenate were loaded in duplicate with decreasing amounts of M. bovis AN5, from an initial concentration of 3 x 105 CFU followed by five successive ten fold dilutions for a final volume of 1ml. This operation was repeated five times over different days.
RESULTS
Colony count of seed lot and disaggregation
The seed lot of M. bovis AN5 had an OD650 of 0.658 corresponding to 3 x 108 CFU/ milliliter (determined by plating and counting after 35 days of development). Three methods were tested to break mycobacterial clumps, use of a Potter homogenizer followed by filtration through 5 µm filter; repeated passages through a tuberculin needle followed by filtration or not. Methods involving filtration yielded little clumping by Ziehl Neelsen staining, but a drastic decrease in the number of mycobacteria. The passage through a needle proved to be simple and suitable for the purpose and this procedure was therefore taken as standard methodology (Figure 1).
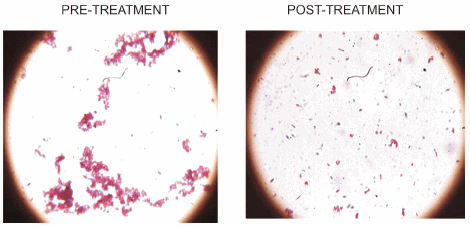
Figure 1. Disaggregation of M. bovis clumps. Ziehl-Neelsen staining was used for observation after the repeated passage of mycobacteria by a tuberculin needle (method c in Materials and Methods). Magnification 40x.
Testing the optimal concentration of PA to conjugate to the beads
Four dilutions of cell extract and culture supernatants were made: 102, 103, 104, and 105. In all but the last supernatant dilution, a positive reaction of the PA with cell extract and culture supernatant was observed by Western blot (Figure 2). Therefore, we concluded that the antiserum was working properly to identify the presence of mycobacteria.

Figure 2. Optimal concentration of polyclonal anti-M. tuberculosis sera to react to M. bovis cell extract and culture supernatants. Western Blot using lyophilized sera was reconstituted and tested in Western blot against a M. bovis AN5 cell extract (E) and culture supernatant (SB). Both materials were subjected to PAGE-SDS and four decreasing dilutions of polyclonal sera were used, 1:100 (1), 1,000 (2), 10,000 (3) and 100,000 (4). MWM, molecular weight marker
To assess the efficiency of the antibody conjugation, two fixed values of 100 and 1,000 CFU were tested varying the antibody concentration. This experiment was conducted in duplicate. IMS-PCR was positive at all the dilutions (Figure 3). Given these results, we decided to use the concentration of 0.7 μg /ml of polyclonal beads, as both tests yielded satisfactory results to both mycobacterial loads (100 and 1,000 CFU). With regard to the optimization of the time required for a better binding of mycobacteria to the beads, no differences were observed between the different incubation times (1, 1.5 and 2 hours). For this reason, we chose 1 h of incubation time.

Figure 3. IMS-PCR to assess optimal polyclonal antibody conjugated to beads. Two bacterial loads were used: 1000 and 100 CFU. Four different dilutions of PA were used: 1 (0.07 μg of PA/μl beads), 2 (0.7 µg/µl), 3 (7 µg/µl) and 4 (15 µg/µl). Amplifications were carried out in duplicate. MWM, molecular weight marker, C+: positive control and C-: negative control
IMS-PCR technique
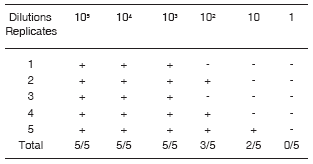
To evaluate the detection limit of the IMS-PCR assay, seven dilutions of the original culture (3 x 108) to 106, 105, 104, 103, 102, 10, and 1 CFU/ml in PBS Tween 0.5% were made. Figure 4 shows that the detection limit of the system was 10 CFU/ml. According to these results, 100% of the reactions had positive results in dilutions containing 105, 104, 103, 102, and 10 CFU, while in the samples containing 1 CFU only 40% of the reactions were positive (2 / 5) ( Table 1 ).

Figure 4. Detection limit of the IMS-PCR assay using bacteria diluted in PBS. MWM, molecular weight marker. C+: positive control and C-: negative control.
Table 1: Detection limit in PBS

IMS-PCR in tissue samples
In a first attempt, the methodology was tested in experimentally contaminated tissue samples, showing only negative reactions except for the tubes inoculated with the highest mycobacterial concentration, as observed in Figure 5A.

Figure 5. Detection limit of the IMS-PCR assay using bacteria diluted in bovine tissues, before (A) and after (B) optimization. MWM, molecular weight marker. C+: positive control and C-: negative control.
Because of these results, we then evaluated the possible presence of PCR inhibitors in the tissue sample. To this end, 900 µl of tissue homogenate were added to each of the dilutions of the M. bovis seed lot (105, 104, 103, 102, 10, 1 CFU). In another condition, a lesser amount of homogenate (100 μl), diluted in 900 µl of PBS 1X, was used to reduce the possible inhibitors present in the tissue. These tubes were contaminated with the same concentrations of mycobacteria as detailed above and showed an increase in the amplification (data not shown), thus indicating a possible partial inhibition. In order to reduce the burden of probable inhibitors, we also carried out successive dilutions of the IMS product (1:2, 1:5, 1:10, 1:20). While some positive results were achieved, these variables were not able to establish a clear cause of the lack of sensitivity. To solve this problem, we decided to include the following changes to the procedure originally proposed:
a) increasing the number of washes of magnetic beads from 3 to 5.
b) increasing the boiling time of post-capture beads from 15 min to 30 min. This step aims to obtain a better lysis and release of mycobacterial DNA.
c) increasing the spin time of the post-capture beads from 5 to 20 min to avoid pipetting magnetic beads with heavy metals that can interfere with the PCR reaction.
By implementing these changes, we observed a significant increase in the positive signals recorded in those samples inoculated with lower concentrations of mycobacteria. Figure 5B and Table 2 show that reactions had positive results in dilutions containing 100,000, 10,000 and 1,000 CFU in 100% of the cases, while the samples that contained 100 and 10 CFU resulted in 60% and 20% positive amplification respectively. At the 1 CFU dilutions, no amplification was observed. As a control, tissue samples were artificially contaminated as previously described and PCR was performed without capture. Under these conditions, only the dilution containing 100,000 CFU resulted positive.
Table 2: Limit of detection in tissues
DISCUSSION
The present work aimed to improve the molecular diagnosis of M. bovis in tissue samples, using immunomagnetic capture of bacteria and subsequent PCR amplification. This system has the advantage of being fast, since it takes only 24 hours. In addition, it is simple to implement and easy to adapt to any laboratory since it does not require sophisticated equipment or expensive reagents. In this way, a single operator can perform 24 samples during the process without major difficulties. The usefulness of this system has been previously demonstrated by Antognoli et al. (1) in experimentally infected milk. Similarly, Romero et al. (15) described that PCR is 28 times more sensitive in the diagnosis of M. tuberculosis complex than traditional culture and direct microscopy. These two works confirm the advantage of PCR in the diagnosis of M. bovis. In addition, unlike Romero et al. (15), who applied the system on samples from TST-positive animals, Zanini et al. (23) described a more effective use of this diagnostic tool applying the system in tissue samples with presence of gross lesions compatible with tuberculosis.
Numerous works have evaluated the use of PCR as a tool for diagnosing mycobacterial infection in various clinical samples. Some have detected M. bovis in milk samples (1-25), whereas others have detected M. bovis directly in bovine tissue (10, 16, 19, 22). Miller et al. (11) demonstrated that PCR is a reliable technique for the identification of M. bovis in tissues embedded in paraffin in which M. bovis could not be cultured. Grant et al. (6) developed IMS-PCR for the detection of M. avium subsp. paratuberculosis in milk samples.
The polyclonal antibody used in the IMS-PCR showed recognition of the mycobacterial extract and supernatant, suggesting a broad recognition of whole mycobacterial cells. We also evaluated the analytical sensitivity of IMS-PCR in PBS infected with decreasing concentrations of mycobacteria. After repeating several trials, we found that the detection limit in this material was 10 CFU/ml in 100% of the samples and that it was 40% efficient when PBS was infected with 1 CFU (Table 1). We thus consider it an acceptable system to be applied to bovine tissues. However, it is important to highlight the following: although the primers used amplified the IS6110 insertion sequence, this segment is present in a small number of copies in the M. bovis genome. In spite of this low copy number, the primers used here provide a high efficiency of target amplification PCR (24). Therefore, we decided to base our work on the positive experiences previously described by other authors (10, 11, 21, 23, 25, 26). Our decision to use IS6110 differs from that made in other studies (19, 22), where the IS1081 insertion sequence present in a higher number of copies (six) in the M. bovis genome was used. In our hands, the sensitivity of IS6110 vs. IS1081 favored the former, thus justifying its use because of its greater potential for amplification of the test (24). We also incorporated the cycling variant known as "touch down", according to that described by Zumarraga et al. (25), who found a two log increase in sensitivity compared with conventional PCR.
Our findings not only show an advantage in the use of IMS but also the possibility to obtain an acceptable detection threshold. It should be emphasized that a detection threshold as low as 10 CFU as that found in PBS-infected tissues, was not reproduced in experimentally infected tissue samples. After a series of tests in tissues, the detection threshold was 1,000 CFU in 100% of the trials. This difference in the detection limit of PBS versus tissues, is probably due to the presence of amplification inhibitors, as only those samples containing more than 1,000 mycobacteria were amplified. Although we could not identify the factors that inhibited the reaction, we demonstrated the need to improve several steps (more washes, a longer boiling time and a longer centrifugation time) in order to avoid false negatives. These results reveal a slight difference in sensitivity as compared to that reported by Wards et al. (22), who described a detection limit in the system between 200 and 500 bacteria in tissues experimentally infected with mycobacterial culture. These values differ from those recently described by Parra et al. (13), who described a detection limit of around 2 to 3 mycobacterial genomes, similar to that described previously (14, 16). Although the experimental sensitivity found by Parra et al. (73.87%) is slightly higher than that described in other studies (around 70%) (10, 19, 23), it is important to point out that the work by Parra et al. (13) was carried out on TB compatible lesion samples, where the sensitivity is directly related to the sampling strategy and the type of injuries.
It is known that mycobacteria present difficulties with their DNA extraction compared to other bacteria or eukaryotic cells, due to the robust cell wall and "clumping". Antognoli et al. (1) mentioned "clumping" as a constraint to correctly estimating the concentration of mycobacteria used to infect, playing an important role in the lack of result reproducibility. For this reason, CFU concentrations should be considered an approximation to the value described. However, the use of the C method (passages through the Tuberculin syringe) to reduce this property proved to be adequate and simple in its application.
This test can provide results in 48 hours, i.e. in a much shorter time than bacteriological culture, where 24-48 days in conventional Stonenbrink and Lowestein Jensen medium are needed, or than the BACTEC radiometric culture radiometric system (BACTEC) described by Negi et al. (12), where 12.8 days are needed or than the BACTEC MGIT 960 System described by Sorlozano et al. (18), where 15.3 days are needed. In this respect IMS-PCR might appear as an alternative. Therefore, the test presented here can be a tool for a national control and eradication campaign of bovine tuberculosis. This technique could result in rapid (48 hours) diagnosis, without the limitation of keeping the viability of the etiologic agent in the sample to be analyzed. To improve the sensitivity of this system and its validation as an alternative tool for diagnosis of bovine tuberculosis, a test of tissue samples from naturally infected animals should be carried out.
Acknowledgements: We thank Pablo Huertas and Liliana Rodriguez from the Institute of Pathobiology (INTA) for their technical assistance. We also thank Martín Zumarraga and Virginia Meikle from the Institute of Biotechnology (INTA) and Elida Gentilini (University of Buenos Aires) for their advice. This work was supported by grants from INTA. AC is a researcher from the National Research Council of Argentina (CONICET).
1. Antognoli MC, Salman MD, Triantis J, Hernández J, Keefe T. A one tube nested PCR for the detection of Mycobacterium bovis in spiked milk samples: an evaluation of different concentration and lytic techniques for M. bovis. J Vet Diagn Invest 2001; 13: 111-6. [ Links ]
2. Aranaz A, Liébana E, Gómez-Mampaso E, Galán JC, Cousins D, Ortega A, et al. Mycobacterium tuberculosis subsp. caprae. subsp. nov. a taxonomic study of a new member of the Mycobacterium tuberculosis complex isolated from goats in Spain. Int J Syst Bacteriol 1999; 49: 1263-73. [ Links ]
3. Cousins DV, Bastida R, Cataldi A, Quse V, Redrobe S, Dow S, et al. Tuberculosis in seals caused by a novel member of the Mycobacterium tuberculosis complex: Mycobacterium pinnipedii sp. nov. Int J Syst Evol Microbiol 2003; 53-5: 1305-14. [ Links ]
4. Di Lonardo M, Isola NC, Ambroggi M, Rybko A. Mycobacteria in HIV-infected patients in Buenos Aires. Tubercle Lung Dis 1995; 76: 185-9. [ Links ]
5. Garnier T, Eiglmeier K, Camus JC, Medina N, Mansoor H, Pryor M, et al. The complete genome sequence of Mycobacterium bovis. Proc Natl Acad Sci USA 2003; 100: 7877-82. [ Links ]
6. Grant IR, Ball HJ, Rowe MT. Isolation of Mycobacterium paratuberculosis from milk by immunomagnetic separation. Appl Enviromnt Microbiol 1998; 64: 3153-8. [ Links ]
7. Haddad N, Masselot M, Durand B. Molecular differentiation of Mycobacterium bovis isolates. Review of main techniques and applications. Res Vet Sci 2004; 76: 1-18. [ Links ]
8. Hermans PW, van Soolingen D, Dale JW, Schuitema AR, McAdam RA, Catty D, et al. Insertion element IS986 from Mycobacterium tuberculosis: a useful tool for diagnosis and epidemiology of tuberculosis. J Clin Microbiol 1990; 28: 2051-8. [ Links ]
9. Khare S, Ficht TA, Santos RL, Romano J, Ficht AR, Zhang S, et al. Rapid and sensitive detection of Mycobacterium avium subsp. paratuberculosis in bovine milk and feces by a combination of immunomagnetic bead separation-conventional PCR and Real-Time PCR. J Clin Microbiol 2003; 42: 1075-81. [ Links ]
10. Liébana E, Aranaz A, Mateos A, Vilafranca M, Gomez-Mampaso E, Tercero JC, et al. Simple and rapid detection of Mycobacterium tuberculosis complex organisms in bovine tissue samples by PCR. J Clin Microbiol 1995; 35: 33-6. [ Links ]
11. Miller JM, Jenny A, Rhyan J, Saari D, Suarez D. Detection of Mycobacterium bovis in formalin-fixed, paraffin-embedded tissues of cattle and elk by PCR amplification an IS6110 sequence specific for Mycobacterium tuberculosis complex organisms. J Vet Diagn Invest 1997; 9: 244-9. [ Links ]
12 . Negi SS, Khan SF, Gupta S, Pasha ST, Khare S, Lal S. Comparison of the conventional diagnostic modalities, Bactec culture and polymerase chain reaction test for diagnosis of tuberculosis. Indian J Med Microbiol 2005; 23: 29-33. [ Links ]
13. Parra A, García N, García A, Lacombe A, Moreno F, Freire F, et al. Development of a molecular diagnostic test applied to experimental abattoir surveillance on bovine tuberculosis. Vet Microbiol 2008; 18; 127: 315-24. [ Links ]
14. Rodriguez JG, Mejia GA, Del Portillo P, Patarroyo ME, Murillo LA. Species-specific identification of Mycobacterium bovis by PCR. Microbiology 1995; 141: 2131-8. [ Links ]
15. Romero RE, Garzón DL, Mejía GA, Monroy W, Patarroyo ME, Murillo LA. Identification of Mycobacterium bovis in bovine clinical samples by PCR species-specific primers. Can J Vet Res 1999; 63: 101-6. [ Links ]
16. Roring S, Hughes MS, Skuce RA, Neill SD. Simultaneous detection and strain differentiation of Mycobacterium bovis directly from bovine tissue specimens by spoligotyping. Vet Microbiol 2000; 70: 227-36. [ Links ]
17. Sequeira MD, Latini OA, López ML. Tuberculosis bovina en seres humanos. II. Período 1977-1989, Rev Arg Tórax 1990; 51: 13-7. [ Links ]
18. Sorlozano A, Soria I, Roman J, Huertas P, Soto MJ, Piedrola G, et al. Comparative evaluation of three culture methods for the isolation of mycobacteria from clinical samples. J Microbiol Biotechnol 2009; 19: 1259-64. [ Links ]
19. Taylor GM, Worth DR, Palmer S, Jahans K, Hewinson RG. Rapid detection of Mycobacterium bovis DNA in cattle lymph nodes with visible lesions using PCR. BMC Vet Res 2007; 3: 1-12. [ Links ]
20. Thierry D, Brisson-Noël A, Vincent-Lévy-Frébault V, Nguyen S, Guesdon JL, Gicquel B. Characterization of a Mycobacterium tuberculosis insertion sequence IS6110 and its application in diagnosis. J Clin Microbiol 1990; 28: 2668-73. [ Links ]
21. Vitale F, Capra G, Maxia L, Reale S, Vesco G, Caracappa S. Detection of Mycobacterium tuberculosis complex in cattle by PCR using milk, lymph nodes, aspirates and nasal swabs. J Clin Microbiol 1998; 36: 1050-5. [ Links ]
22. Wards BJ, Collins DM, de Lisle GW. Detection of Mycobacterium bovis in tissues by polymerase chain reaction. Vet Microbiol 1994; 43: 227-40. [ Links ]
23. Zanini MS, Moreira EC, Lopes MT, Oliveira RS, Leão SC, Fioravanti RL, et al. Mycobacterium bovis: Polymerase chain reaction identification in bovine lymph node biopsies and genotyping in isolates from southeast Brazil by spolygotyping and restriction fragment length polymorphism. Mem Inst Oswaldo Cruz 2001; 96: 1-5. [ Links ]
24. Zumarraga MJ, Cataldi A, Bigi F, Alito A, Romano MI. Aplicación de la reacción en cadena de la polimerasa (PCR) en la detección de micobacterias en leche. Rev Arg Microbiol 1999; 31 (Supl.1): 4-5. [ Links ]
25. Zumarraga MJ, Meikle V, Bernardelli A, Abdala A, Tarabla H, Romano MI, et al. Use of touch-down polymerase chain reaction to enhance the sensitivity of Mycobacterium bovis detection. J Vet Diagn Invest 2005; 17: 232-8. [ Links ]
26. Zumarraga MJ, Paolicchi F, Garbaccio S, Giofree A, Cataldi A. Aplicación de la PCR en la detección de Mycobacterium bovis en muestras de tejido de terneros. Vet Arg 2001; 18: 668-76. [ Links ]
Recibido: 18/05/2010
Aceptado: 09/08/2010














