Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Revista argentina de microbiología
Print version ISSN 0325-7541
Rev. argent. microbiol. vol.44 no.2 Ciudad Autónoma de Buenos Aires Apr./June 2012
INFORME BREVE
Detection of the ompA gene of Chlamydophila pecorum in captive birds in Argentina
María C. Frutos1*, Fernando Venezuela1, Ximena Kiguen1, Viviana Ré1, Cecilia Cuffini1
1Instituto de Virología "Dr. J. M. Vanella", Facultad de Ciencias Médicas, Universidad Nacional de Córdoba. Enfermera Gordillo Gómez s/n Ciudad Universitaria (5016) Córdoba, Argentina.
*Correspondence. E-mail: mariaceliafrutos@gmail.com
ABSTRACT
Bacteria belonging to the family Chlamydiaceae cause a broad spectrum of diseases in a wide range of hosts, Including humans, other mammals and birds. However, very little is known about chlamydial infections in birds in our region. In the present study, we examined 28 clinically normal birds In illegal captivity that were confiscated in the province of Córdoba, Argentina. The objective was to detect Chlamydophila spp. in cloacal swabs by genetic analysis of the ompA gene. Nested-PCR of the ompA gene identified five samples as Chlamydophila pecorum and the sequence analysis demonstrated the presence of the ompA gene of C. pecorum In these birds. On the other hand, Chlamydophila psittaci was not detected. These birds could be either asymptomatic reservoirs or subclinical carriers of C. pecorum. This is the first report of the detection of C. pecorum in Argentina.
Key words: Chlamydophila pecorum; Birds In captivity; OmpA gene; Sequence analysis
RESUMEN
Detección del gen ompA de Chlamydophila pecorum en aves en cautiverio, en la Argentina.
Las bacterias que pertenecen a la familia Chlamydiaceae causan un extenso espectro de enfermedades en una amplia gama de huéspedes, incluidos los seres humanos, otros mamíferos y aves. Sin embargo, se sabe muy poco acerca de las infecciones por clamidias en aves de nuestra reglón. Esta Investigación examinó 28 aves clínicamente normales mantenidas en cautiverio ¡legal, que fueron confiscadas en Córdoba, Argentina. El objetivo fue detectar Chlamydophila spp. en hisopados cloacales por análisis del gen ompA. La PCR anidada del gen ompA reveló la presencia de Chlamydophila pecorum en cinco muestras. El análisis de secuencias demostró la presencia del gen ompA de C. pecorum en estas aves. Por el contrario, Chlamydophila psittaci no se detectó. Estas aves pueden ser reservónos asintomáticos o portadores subclínlcos de C. pecorum. Este es el primer informe de la detección de C. pecorum en la Argentina.
Palabras clave: Chlamydophila pecorum; Aves en cautiverio; Gen ompA; Análisis de secuencias
Chlamydophila psittaci, the causing agent of avian chlamydiosis, occurs worldwide and has been detected in a wide variety of both wild and domestic birds. However, other clamydiae also have a zoonotic potential (3).
Chlamydophila pecorum strains have been isolated from ruminants, swine and koalas in several countries. C. pecorum is associated with conjunctivitis, encephalomyelitis, enteritis, pneumonia, polyarthritis, abortion, and reproductive and urinary tract diseases (1,3,8). In an assay carried out in free healthy pigeons in Japan, three fecal samples were found to be C. pecorum-positive by PCR (7).
The epidemiology of Chlamydia infection in animals in Argentina is unknown. Thus, the aim of the present study was to detect Chlamydia spp. in illegally captive birds in Córdoba city, Argentina.
Cloacal swabs were collected from 28 birds living in illegal captivity without any clinical signs or evidence of chlamydiosis and were referred to the Instituto de Virología, Facultad de Ciencias Médicas, Universidad Nacional de Córdoba, Argentina. According to provincial law N.° 1751, which prohibits catching and commercialization, these birds were considered illegal possessions. Therefore, they were confiscated by the Secretaría Provincial del Medio Ambiente in September 2010. The cotton swabs were placed in 1 ml of sucrose-phosphate-glutamate, and 200 pi were subjected to DNA extraction using the Accuprep Genomic DNA Extraction Kit (BIONEER, Alameda, CA, USA) according to the manufacturer's instructions.
DNA extract (5 ul) was used to amplify a 576-bp fragment of the variable domains II, III and IV of the ompA gene of Chlamydophila, using primers 191 CHOMP (GCI YTI TGG GAR TGY GGI TGY GCI AC) and CHOMP 371 (TTA GAA ICK GAA TTG IGC RTT IAY GTG IGC IGC), as described by Kaltenbock et al. (2) and modified by Sachse and Hotzel (5). Five samples were positive by PCR.
Nested-PCR using primers 218PSITT (GTA ATT TCI AGC CCA GCA CAATTY GTG)/CHOMP 336 (CCR CAA GMT TTT CTR GAY TTC AWY TTG TTR AT) showed that none of the samples was positive for C. psittaci (389 bp) and that five (17.85 %) were positive for C. pecorum (441 bp) with primers 204 PECOR (CCA ATA YGC ACA ATC KAA ACC TCG C)/CHOMP 336(CCR CAA GMT TTT CTR GAY TTC AWY TTG TTR AT). The latter were confirmed by sequencing.
For sequence analysis, the nested-PCR products were purified with the QIAquick Gel Extraction Kit (Qiagen, Valencia, CA, USA) and subjected to direct nucleotide sequencing reaction in both directions using an ABI automatic sequencer. The ompA sequences obtained were submitted to GenBankand relatedness of newly characterized sequences was assessed by analysis with 2.2.19 Basic Local Alignment Search Tool (BLAST). Phylogenetic analyses were constructed with Mega software version 4, using the Neighbor-Joining method and their reliability was assessed by bootstrap (2,000 replicates) (6). Nine reference sequences of the ompA region of the C. pecorum genome (403 bp) were included in this analysis. The corresponding accession numbers, country of origin, host and clinical condition of these isolates are shown in Table I. The tree was rooted with the ompA sequence of the Chlamydophila abortus strain (accession number CR848038.1).
Table 1. Characteristics of C. pecorum isolates analyzed or used as reference strains

The phylogenetic analysis showed that the five sequences clustered with C. pecorum.
The ompA sequence of the C. pecorum strains was highly homologous and shared more than 98 % similarity with each other. Blast searches revealed that all novel ompA gene sequences shared high homology with the ompA from the iC2, iC3, iC4, 3638/ 3 and 4283/3 strains.
All local strains were characterized by a thymine at position 132, resulting in Asn instead of Lys in the VD IV of the ompA gene (Figure 1).The discrepancy found in this position could be defined as a marker of local strains (Figure 1).

Figure 1. Comparison of MOMP VD IV sequences. Dots represent residues that are Identical to those In IC2, IC3, IC4, 3638/3, and 4283/3 MOMP. The first and the last residues In the IV domain are given as Its position In the complete alignment.
In 98 % of the cases, the bootstrap values in the ompA phylogenetic tree supported the conclusion that all local strains belong to the same group (Figure 2).
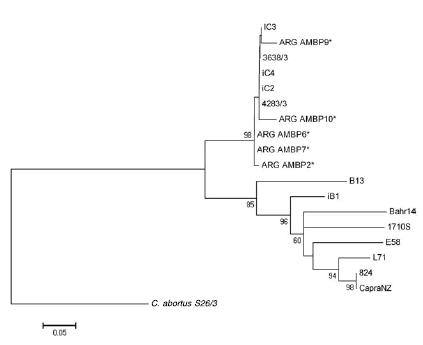
Figure 2. Phylogenetic three analysis of the ompA region of the C. pecorum genome (403 bp) of the Argentinean isolates and C. abortus as rooted. Strains belonging to this study are initiated by ARG and marked with asterisks. The bootstrap (> 50 %) is Indicated In the branch nodes.
This chlamydia has generally been associated with the detection in ruminants (10). However, Tanaka et al. detected this chlamydia in birds similar to those studied here. Nevertheless, the homology with these sequences could not be assessed because they are not available at GenBank.
C. pecorum infection did not appear to be associated with any clinical signs of these birds. Thus, these birds could be either asymptomatic reservoirs or subclinical carriers of C. pecorum.
The epidemiological significance of C. pecorum infection is not clear to us at this stage. Further studies are needed to estimate the zoonotic role of this pathogen.
Competing interests
The authors declare that they have no competing interests.
Acknowledgements: this work was partially supported by SECyT-UNC 05/H181, Mincyt-Cba 1427/09, Mincyt-PId 113/11. This short communication has been prepared In the context of the collaboration promoted by the Provincial Secretariat for Environmental Protection and Sustainable Development, Córdoba, Argentina.
1. Jackson M, White N, Giffard P, Timms P. Epizootiology of Chlamydia Infections in two free-range koala populations. Vet Microbiol 1999; 65: 255-64. [ Links ]
2. Kaltenböeck B, Schmeer N, Schneider R. Evidence for numerous omp1 alleles of porcine Chlamydia trachomatis and novel chlamydial species obtained by PCR. J Clin Microbiol 1997; 35: 1835-41. [ Links ]
3. Longbottom D, Coulter L. Animal chlamydiosis and zoonotic implications. J Comp Pathol 2003; 128: 217-44. [ Links ]
4. NaroskyT, Yzurieta D. Fam. Emberizidae. In: Vázquez Mazzini, editores. Guía para la identificación de aves de Argentina y de Uruguay, 15a edición. Buenos Aires, Argentina, Vázquez Mazzini, 2007, p. 265-75.
5. Sachse K, Hotzel H. Detection and differentiation of Chlamydiae by nested-PCR. Methods Mol Biol 2002; 216: 123-36. [ Links ]
6. Tamura K, Dudley J, Nel M, Kumar S. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 2007; 24: 1596-9. [ Links ]
7. Tanaka C, MiyazawaT, Wataral M, Ishiguro N. Bacteriological survey of feces from feral pigeons in Japan, J Vet Med Sci 2005; 67: 951-3.
8. Yousef Mohamad K, Rodolakis A. Recent advances in the understanding of Chlamydophila pecorum infections, sixteen years after It was named as the fourth species of the Chlamydiaceae family. Vet Res 2010; 41: 27-37. [ Links ]
Recibido:30/8/2011- Aceptado: 4/5/2012














