Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Revista argentina de microbiología
Print version ISSN 0325-7541
Rev. argent. microbiol. vol.44 no.2 Ciudad Autónoma de Buenos Aires Apr./June 2012
ARTÍCULO ORIGINAL
Biodegradation of phenol in static cultures by Penicillium chrysogenum ERK1: catalytic abilities and residual phototoxicity
Erika A. Wolskm*,Viviana Barrera2, Claudia Castellari3, Jorge F. González1
1Facultad de Ingeniería, Universidad Nacional de Mar del Plata, J. B. Justo 4302 (7600) Mar del Plata;
2IMYZA, CNIA, INTA-Castelar, Castelar; "Microbiología de Suelos y Alimentos, Unidad Integrada EEA INTA-Balcarce, Facultad de Ciencias Agrarias, Ruta Nacional 226 Km 73,5; Balcarce. Argentina. Correspondence. E-mail: ewolski@mdp.edu.ar
ABSTRACT
A phenol-degrading fungus was isolated from crop soils. Molecular characterization (using internal transcribed spacer, translation elongation factor and beta-tubulin gene sequences) and biochemical characterization allowed to identify the fungal strain as Penicillium chrysogenum Thorn ERK1. Phenol degradation was tested at 25 °C under resting mycelium conditions at 6, 30, 60, 200, 350 and 400 mg/l of phenol as the only source of carbon and energy. The time required for complete phenol degradation increased at different initial phenol concentrations. Maximum specific degradation rate (0.89978 mg of phenol/day/mg of dry weight) was obtained at 200 mg/l. Biomass yield decreased at initial phenol concentrations above 60 mg/l. Catechol was identified as an intermediate metabolite by HPLC analysis and catechol dioxygenase activity was detected in plate assays, suggesting that phenol metabolism could occur via ortho fission of catechol. Wheat seeds were used as phototoxicity indicators of phenol degradation products. It was found that these products were not phytotoxic for wheat but highly phytotoxic for phenol. The high specific degradation rates obtained under resting mycelium conditions are considered relevant for practical applications of this fungus in soil decontamination processes.
Key words: Penicillium chrysogenum; Soil fungus; Phenol; Biodegradation; Phytotoxicity
RESUMEN
Biodegradación de fenol en cultivos estáticos por Penicillium chrysogenum ERK1 : habilidades catalíticas y fitotoxicidad residual. Un aislamiento fúngico capaz de degradar fenol como única fuente de carbono y energía fue aislado de suelos agrícolas. La caracterización molecular (basada en el empleo de secuencias de espaciadores de transcriptos internos, de factores de la elongación de la traducción y del gen de la beta-tubulina) y la caracterización bioquímica permitieron identificar a esta cepa como Penicillium chrysogenum Thom ERK1. Se estudió la degradación de fenol a 25 °C en cultivos estáticos con 6, 30, 60, 200, 350 y 400 mg/l de fenol inicial. El tiempo requerido para completar la degradación de fenol aumentó al elevarse las concentraciones iniciales de dicho compuesto. La máxima tasa de degradación específica (0,89978 mg de fenol/día/mg de peso seco) se obtuvo con 200 mg/l. El rendimiento en biomasa disminuyó con concentraciones Iniciales de fenol mayores de 60 mg/l. Se identificó al catecol como intermediarlo metabolico por HPLC y se observó actividad de catecol dioxigenasa en placa, lo que sugiere que el metabolismo de degradación del fenol ocurre vía orto fisión del catecol. Se utilizaron semillas de trigo como indicadores de fitotoxicidad de los productos de degradación. Estos productos no fueron fitotóxicos para trigo, mientras que el fenol mostró una alta fitotoxicidad. La alta tasa de degradación específica obtenida en condiciones estáticas resulta de gran interés para la aplicación de este hongo en procesos de descontaminación de suelos.
Palabras clave: Penicillium chrysogenum; Hongos del suelo; Fenol; Biodegradación; Fitotoxicidad
INTRODUCTION
Soil and water contamination is now considered a serious problem in many industrialized countries. Phenol contamination may arise from a variety of sources of industrial wastewater, such as those from coal refineries, phenol manufacturing pharmaceuticals, industries of resins, paints, dyes, petrochemicals, textiles, pulp and paper mill (9).These phenolic effluents are being discharged into water bodies and this water is used for agriculture and other purposes. Reports of incidents on phenol contamination in the area are scarce. In 2004, some information sources reported that the Iguazu River (Argentina) was affected by the discharges of petrochemical effluents which contain 0.14 mg/l of phenol and other compounds like manganese and aluminium. For these reasons, wastewaters containing phenol and phenolic compounds need an appropriate treatment before discharging them into the receiving water bodies (20). Many physico-chemical techniques are available to degrade these pollutants before discharging them (10). Some of these techniques are effective, but most of them are expensive and may lead to the formation of secondary toxic materials or lower mineralization, or need severe operating conditions (10). For these reasons, biological degradation is a viable and economic alternative, which leads to the complete mineralization of the xenobiotic.
On the other hand, bioremediation is a well known treatment for soil contamination, which employs the use of microorganisms that are either naturally occurring or introduced into the soil in order to degrade pollutants (13). Several fungal strains have been reported to degrade phenol as the only source of carbon and energy (5, 12, 18, 19, 24), but many of them are phytopathogens or have high nutritional requirements or fail to colonize the soil and are difficult to apply for soil bioremediation. For these reasons, there is interest in the study of new, non-pathogenic soil fungal isolates.
Pénicillium species are commonly found in food, indoor air and soils. Particularly, the Pénicillium chrysogenum has been found on dried cereals, salted meat and many other low water activity foods, but is also common in indoor air environments and salty soils (22). Several members of the genus Pénicillium are good hydrocarbon-assimilating organisms and there are many reports showing their ability to transform xenobiotic compounds into less mutagenic products (4, 10). There are many examples of that: Pénicillium simplicissimun SK9117 was able to grow in 800 mg/l of phenol (5), P. chrysogenum CLONA 2 isolate also completely degraded 300 mg/l of phenol in the presence of sodium chloride (58.5 g/l) (4, 9). Most of these studies were carried out in fermentors under intensive stirring. These conditions do not necessarily reflect those prevalent in an actual soil decontamination process.
In this work, we report the isolation and both molecular and biochemical identification of a P. chrysogenum strain called ERK1, isolated from crop soils in Argentina. This fungus was able to degrade phenol as the only source of carbon and energy with high degradation rates under resting mycelium conditions. This is considered relevant for practical applications in soil decontamination processes where the fungus is also in static conditions. Degradation potential, metabolic intermediate and phytotoxicity assays were carried out.
MATERIALS AND METHODS
Chemicals
All the reagents used during this study were of analytical grade, except for phenol and catechol that were of chromatographic grade (purity 99 %) from Sigma-Aldrich (St. Louis, USA). HPLC-grade acetonitrile was from Sintorgan (Argentina).
Microorganism isolation and growth conditions
Soil samples were collected from commercial crop soils from Balcarce, Buenos Aires province, Argentina. The soil samples (3 g weight) were mixed with 10 ml sterile water. Serial dilutions of the culture were prepared and spread on mineral medium agar plates supplemented only with 25 mg/l of phenol as a carbon source. The plates were incubated at room temperature for 3 days. The suspected type of colony was purified and was maintained in potato dextrose agar (PDA, Gibco) at room temperature for 14 days (without phenol).
For the degradation assays, the fungus was inoculated directly from the PDA plate into 150 ml of liquid minimal salt medium (LMS) supplemented with different concentrations of phenol as the only source of carbon and energy. The LMS contained: deionized water 1000 ml, MgS04-7H20 0.1 g, K2HP04 0.1 g, NH4N031 g, KCI 0.1 g, and 25 ul of trace element solution (in mg/l: MnS04 15.4, FeCI3 40, ZnS04.7H20 6.3, CuS04.5H20 2.5, NH4.M07.02.4H20 0.5). pH was adjusted to 6.0.
The selected isolates were identified by physiological, biochemical and molecular tests.
Morphological characteristics
The fungus was inoculated onto different culture media: Malt Extract Agar (MEA, Britania, Argentina), Czapek Yeast extract Agar (CYA, Britania, Argentina) and 25 % Glycerol Nitrate (G25N, Britania, Argentina) to observe the different morphological characteristics (colony diameter, color, pigments, exudates, etc.) for its identification according to Pitt (17), Pitt and Hocking (16) and Samson et al. (21).
Antibiotic production
In order to confirm that the fungus was a Penicillium chrysogenum isolate, its ability to produce B-lactam antibiotics was analyzed by a diffusion bioassay using Micrococcus luteus (ATCC 9341) as described by Castellan et al. (1 ),since the diameter of the growth inhibition zone is characteristic for each species and is also used as a complement to identify the fungal isolate (1, 8).
The presence of B-lactam antibiotic was also analyzed in the culture medium were P. chrysogenum was grown with phenol as the only source of carbon and energy.
Nuclear number per cell
To observe the number of nuclei per cell, small portions of mycelia previously grown on PDA in darkness for 24 h at 25 °C were submerged in 0.01 % acridine orange aqueous solution during 10 seconds. The method applied is a modification from theYamamoto and Uchida's staining method (28). The stained mycelium was observed under epifluorescent light with an OLYMPUS BX 51 microscope. Digital photographs were taken using the Cool Snap-Pro System. The number of nuclei was counted in 20 cells.
Ehrlich test
The fungal isolate was examined for production of cyclopiazonic acid and other alkaloids reacting with Ehrlich' reagent using a filter paper method as described by Frisvad and Samson (2).
DNA extraction
The ERK1 strain was grown on potato dextrose broth (PDB) for 3 days. The mycelium obtained was dried, freezed and disrupted with a hand-operated pellet pestle and DNA was obtained with DNeasy Plant Mini Kit QIAGEN according to the manufacturers' protocol. The resuspended DNA was stored at -20 °C. The DNA was quantified by electrophoresis gel with 0.8 % agarose and NanodropT 2000 (Thermo Scientific).
Polymerase Chain Reaction
Internal transcribed spacer (ITS). Genomic DNA was amplified with primers ITS1 (5'-TCCGTAGGTGAACCTGCGG-3') and ITS4 (5'-TCCTCCGCTTATTGATATGC-3') to amplify the region ITS1-5.8s-ITS2 (33). Amplifications were performed using 20 mM Tris-HCI (pH 8.4), 50 mM KCI, 0.2 mM dNTPs, 0.2uM primer, 3 mM MgCI2, 0.4 U Taq Invitrogen (Brazil) with 10-100 ng genomic DNA in a 50 ul total volume. The amplification program was as follows: 1 min at 94 °C, 1 cycle; 15 s at 94 °C, 15 s at 58 °C, 15 s at 72 °C, 30 cycles; 7 min at 72 °C, 1 cycle.
Beta-tubulin. To amplify the beta-tubulin gene, primer pairs Bt2a (5'- GGTAACCAAATCGGTGCTGCTTTC-3') and Bt2b (5'- ACCCTCAGTGTAGTGACCCTTGGC-3') (3) were applied. Amplifications were performed with the same conditions above with the amplification program reported in reference 22.
Translation elongation factor (TEF). The amplification of TEF sequences was performed using the primers/pair Ef728M (5'-CATYGAGAAGTTCGAGAAGG-3') and Ef2 (5'-GGARGTACCAGTSATCATGTT-3') following the procedure in Samuels and Ismaiel (23).
Sequencing
The amplification products were cleaned with the Wizard® SV Gel and PCR Clean-Up System (PROMEGA) and visualized with gel electrophoresis using 1 % agarose stained with ethidium bromide. The PCR products were sequenced in both directions under BigDyeTM Terminator v 3.1 (Applied Biosystems) based on the Sanger's method. The reacted products were purified using ethanol precipitation and run with Genetic Analyzer 3130xlatUGB, Unidad de Genomica, Instituto de Biotecnologia, INTA). Contigs were assembled using the CAP3 Sequence Assembly Program (PBIL, France) (6).
Phylogenetic Analysis
Alignments of the partial beta-tubulin gene sequences were constructed automatically using a built-in CLUSTALW implementation in MEGA software version 4 (27).
Phylogenetic and molecular evolutionary analyses were conducted using MEGA 4.1. All positions containing gaps and missing data were eliminated from the dataset (Complete Deletion option). The evolutionary history was inferred using the Maximum Parsimony method; bootstrap analysis was performed in 1000 replicates with random sequence addition (10 replicates) to estimate branch support. The MP tree was obtained using the Close-Neighbor-Interchange algorithm with search level 3 in which the initial trees were obtained with the random addition of sequences (10 replicates). Statistics including tree length (L), consistency index (CI), retention index (Rl) and composite index were also calculated. The sequences of the beta-tubulin gene and rDNA-ITS obtained were submitted to GenBank, accession numbers HQ336382 and HQ336383. Thirteen sequences from the GenBank database were used to construct the phylogenetic analyses (Fig. 3); the sequences were chosen to represent sections of Pénicillium related with the ERK1 strain, following the classification of Samson et al. (22). Eupenicillium brefeldianum was applied as outgroup taxa.
Phenol degradation
Degradation of phenol was analized in submerged cultures in 250 ml flasks with 150 ml of LMS medium supplemented at different phenol concentrations as the only source of carbon. Each flask was inoculated with 4 PDA agar discs containing the fungal mycelium. The cultures were incubated at room temperature in the dark in order to avoid phenol destruction under resting mycelium conditions.
Non-inoculated flasks with LMS supplemented with phenol were used as controls.
At different times after inoculation the mycelium from each flask was filtered and the dry weight was determined. Phenol content of the liquid medium was measured in the filtrates after removal of the mycelia. All experiments were carried out in triplicate. Results show the mean value of three independent experiments.
Specific degradation rates were calculated as follows:

Where: P is phenol concentration in mg/l, t is time in days, 8 is biomass in mg of dry weight per litre of reactor volume.
Dry weight
Dry weight was determined by filtering the mycelium through a Whatman GF/A filter, rinsing twice with distilled water and drying at 100 °C until constant weight. Biomass was calculated as mg of dry weight per volume reactor (I).
Analytical methods
The concentration of phenol was measured using a Waters HPLC system (Millipore, Waters Division, Milford, Massachusetts, USA) consisting of a Model 590 pump, equipped with a UV detector Model 484 variable-wavelength detector set at 270 nm. A computer equipped with HPLC System Manager Software for windows CSW 32 v.1.4 (2002 DataApex Ltd. Czech Republic) was used to acquire and process chromatographic data. The separation was achieved with a Water Spherisorb ODS2 C18 (5 urn) 4.6 x 250 mm analytical column Millipore Corporation, Milford, Massachusetts, USA. Deionized water: acetonitrile (70:30, vol/vol) isocratic system was used as solvent and the flow rate was maintained at one ml/min. These compounds were identified by comparing their retention time with those of similarly treated external standards and by co-chromatography. Under the above conditions, the retention time of phenol and catechol were 10.09 and 5.57, respectively.
Plate assays to assess enzymatic activity
Qualitative assays were performed on agar plates to determine the enzymatic activity involved in phenol degradation. Laccase, peroxidase (manganese peroxidase and lignin peroxidase) and catechol dioxigenase activities were determined as described by Rubilar-Araneda (19), Levin et al. (11) and Shiftman and Cohen (26), respectively.
Phytotoxicity studies
The toxicity of the original and the degraded phenol was assessed by measuring the phytotoxicity effect of LMS, LMS supplemented with phenol 400 mg/l and the residue of phenol degradation on seed germination of wheat (Triticum aestivum), according to Osma et al. (15). Five replicates of 10 seeds were used for each treatment. Germination index (Gl) was calculated as follows: GI=GP x La/Lc, where GP is the number of germinated seeds expressed as a percentage of control values (LMS). La is the average value of root length in the phenol solutions and Lc is the average value of root length in the control.
RESULTS
Identification of the fungal isolate
The fungus was grown on MEA, CYA and G25N. Mycelia showed white obverse color in all the media tested (Figure 1). The reverse colors varied from white to yellow. In CYA at 25 °C, the fungus showed yellow exudates and colonies were radially sulcated (Figure 1).
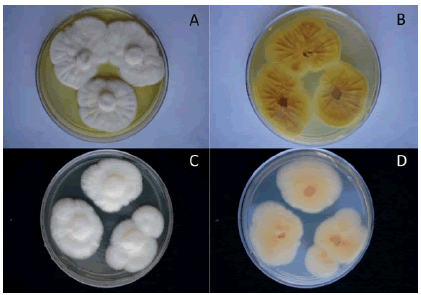
Figure 1. Growth of the fungus on CYA and MEA. A: CYA, 7 days, 25 °C. B: reverse CYA, 25 °C. C: MEA, 7 days, 25 °C. D: reverse MEA, 25 °C.
The fungus grown in MEA medium did not show green color or penicillia at 25 °C. However, when these plates were stored at 4 °C for one month, green color and penicillia were observed (Figure 2 A). Microscopically, penicillia are terverticillate, smooth walled, the conidia width was 3.75 urn and phialide length 12.3 urn (Figure 2 B). The nuclear number per cell was 1 nucleus in hyphal tips and 2-8 nuclei in mature hyphae.

Figure 2. Biochemical characteristics of the isolate. A: colony growth (one month old) in MEA. B: Penicillin 1000X. CAntibiotic-susceptibility test with Micrococcus luteus (ATCC 9341) (susceptible to B-lactam antibiotics). Clear zones indicate growth inhibition around the disk containing the fungal isolate.
The fungus also showed inhibitory activity against Micrococcus luteus (ATCC 9341), producing a growth inhibition zone with a diameter of 45 mm in diffusion bioassays, suggesting the production of B-lactam antibiotics (Figure 2 C).
The isolate was observed for production of cyclopiazonic acid and other alkaloids by the Ehrlich's test, but no reaction was observed.
Taxonomic identification of the fungus based on Pitt and Hocking (16) shows that the fungus isolated in this work is a Pénicillium strain.
Genomic DNA obtained with the extraction procedure yielded 35 ng/ul. A fragment of 600 bp in size was obtained with ITS1/4 primer pair. A sequence of 441 bp was obtained from the contig assembly of the sequences in both directions. The comparison with Megablast showed 100 % homology (E value 0.0) with P. chrysogenum and Pénicillium comune. A fragment of 455 bp in size was obtained from the amplification with pair primers Bt2a/Bt2b. The TEF amplification product was over 500 bp in size, although the corresponding sequence obtained was 265 bp in size due to technical difficulties with the pair primers tested; therefore, it was rejected for further analyses.
From the Parsimony Analysis of the beta-tubulin gene sequences, 4 most parsimonious trees with L=242 steps were obtained, Cl=0.643836, Rl=0.603053 and Composite lndex=0.473472. Branches corresponding to partitions reproduced in less than 50 % bootstrap replicates are collapsed. The percentage of replicate trees in which the a ssociated taxa clustered together in the bootstrap test is shown next to the branches. The most parsimonious assignments of ancestral states for site #1 are shown next to each node. There were a total of 359 positions in the final dataset, out of which 64 were parsimony informative (Figure 3). The clades corresponding to the sections showed high bootstrap values, with 70 % for section Chrysogena, 99 % for Roquefortii and 74 % for Viridicata with the exception of Pénicillium with 27 %.

Figure 3. Phylogenetic tree. Maximum parsimony consensus tree with beta-tubulin gene sequences of type strains of Penicillium taxa. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches. Eupenicillium brefeldianum NRRL 710 was applied as an out group. The bars at the right of the cladogram indicate the sections of subgenus Penicillium classified by Samson et al. (2004).
Phenol degradation
For kinetic studies, the fungus was grown in LMS medium supplemented with phenol as the only source of carbon and energy. Phenol degradation was tested at nominal values of 6,30,60, 200, 350, 400 mg/l under resting mycelium conditions (Figure 4). The actual values of phenol concentrations measured by HPLC were 6.28, 31, 60.9, 200, 347 and 400 mg/l. The incubation of P. chrysogenum ERK1 in the medium containing phenol as the sole carbon source resulted In an Increase In the dry weight of the mycelia over the time, concomitant with the decrease in the concentration of phenol In the culture medium (Figure 4). In the case of 60, 200, 350 and 400 mg/l of phenol, the blomass increases until reaching a plateau value which means that the fungal growth has reached the stationary phase (Figure 4). Biomass yield was calculated for each initial phenol concentration and it decreases with the increase of the initial concentration of phenol until 200 mg/l, after which it remained essentially constant (Table 1).

Figure 4. Growth and biodégradation of phenol by P. chrysogenum in submerged culture. Degradation of phenol was analyzed by HPLC. LMS medium was supplemented at different phenol concentration as the only source of carbon and energy and inoculated with P. chrysogenum. The cultures were incubated in the dark at 25 °C in order to avoid photodestruction of phenol and under resting mycelium conditions. Noninoculated flasks with LMS supplemented with phenol were used as controls. A, B, C, D, E and F corresponding to 6,30,60,200,350 and 400 mg/l of initial phenol concentrations, respectively.
Table 1. Kinetic and yield values as a function of substrate concentration

No further dry weight increase was observed following phenol depletion, which demonstrated the ability of the organism to grow on phenol as the only source of carbon.
The time required for complete degradation of the phenol varied at different initial phenol concentrations (Figure 4). Degradation took longer at higher phenol concentrations. For example, on day 6 the degradation percentages were 100 %, 75 %, 51 %, 25.7 %, 26 % and 8 % for 6, 30, 60, 200, 350 and 400 mg/l, respectively. For the Initial phenol concentration of 200 and 350 mg/l, the conversion percentages were similar. These results agree with the degradation rates observed for both initial phenol concentration (Table 1). Figure 4 also shows that while the biomass remains constant, the degradation of phenol continues.
Specific degradation rates increase until 200 mg/l, where a maximum is observed, and then decrease showing an inhibitory effect (Table 1). These results agree with that observed for biomass yield.
The inhibitory activity against M. luteus (ATCC 9341) by P. chrysogenum was analyzed when phenol was used as a carbon source. In these experimental conditions, the fungus did not inhibit M. luteus (ATCC 9341) growth, suggesting that P. chrysogenum did not produce B-lactam antibiotic when phenol Is used as carbon source.
HPLC analysis showed that phenol was completely degraded when used at an Initial concentration of 200 mg/l (Figure 5 B). Chromatographic profiles revealed the presence of phenol metabolites formed by the fungus at very low concentrations (less than one mg/l) during the kinetic assay (on day 18) (Figure 5 C). One of the major detected compounds had the same retention time (rt) as the external standard catechol. Other products of phenol degradation were found to be minor, but could not be identified structurally. These minor compounds showed peaks around rt: 5.9, 3.29, 3.77 and 3.9, respectively.

Figure 5. HPLC-chromatogram of phenol biodégradation by P. chrysogenum. Kinetics at 200 mg/l of initial phenol concentration was used for this analysis. A: time zero showing the retention time of phenol (Phe: 10.01 min). B: biodégradation of phenol aft er 20 days. C: metabolite formed during the kinetic assay on day 18 after inoculation time.
Neither laccase nor peroxidase activities (Mn peroxidases and Li peroxidases) were detected. However, catechol dioxigenase activity was observed in qualitative assays (data not shown). Fungal mycelium developed a yellow color on the plate sprayed with catechol.
Phytotoxicity assay
The results of phytotoxicity showed that seed germination was 100 % for wheat in the LMS medium used as a control, 95 % for seeds treated with the products of phenol degradation and 6.25 % for 400 mg/l of phenol. No root elongation was observed in the latter case. Seeds treated with phenol degradation products and control seeds showed similar root elongation, about 14 ± 7.1 mm. Therefore, Gl values were: 100 % for both, while seeds treated with 400 mg/l of phenol showed a Gl value of 0.4 %.
DISCUSSION
Taxonomic Identification of the fungus based on Pitt and Hocking (16) shows that the fungus isolated in this work Is a Penicillium strain, which produces a B-lactam antibiotic showing an inhibition zone in diffusion bioassays lacking phenol, typical of P. chrysogenum, as described by Castellari et al. (1).
The phylogenetlc analysis based on beta-tubulin gene sequences grouped the species belonging to the same sections together as expected by the comparison with the parsimony analysis performed by Samson et al. (22), although the topology of the tree obtained showed differences with Section Chrysogena as poliphyletic. ERK1 strain grouped with 93 % bootstrap with P. chrysogenum CBS 306.48. These results are consistent with the high homology observed when analyzed in Megablast with ITS sequences. Although the ITS sequences also yielded high homology with P. commune, the sequence analyses of the beta-tubulin gene allowed to differentiate it in a separate cluster from P. chrysogenum. The relevance of Samson et al's phylogenetlc analysis (22) (Ibid.) lies in that the authors supported the phylogenetic clades based not only on DNA characters but also on phenotypic characters (morphology and cultural characters).
Therefore, from the agreement in molecular and biochemical characterization, it has been demonstrated that the fungal Isolate is very closely related to P. chrysogenum.
There are many studies describing the biodégradation of phenol using Pénicillium Isolates (5, 9, 12, 24, 25). Scow et al. (25) have described the mineralization of phenol at low Initial concentrations by a non-identified Pénicillium species. On the other hand, Pénicillium frequentans Bi 7/2 and P. simplicissimum SK9117 also use phenol as the only source of carbon and energy (5, 12). P. chysogenum CLONA 2 isolate also completely degrades 300 mg/l of phenol in the presence of sodium chloride (58.5 g/l) (9). Moreover, In this work characterization and degradation potential of a new P. chrysogenum ERK1 strain was studied. This fungus shows high specific degradation rates under resting mycelium conditions; which makes it attractive for practical applications in soil decontamination processes. In addition, preliminary results showed that the fungus was able to grow and degrade phenol in artificially contaminated soils (Data not shown).
The specific degradation rates values obtained showed that an inhibition effect occurs. The comparison of the specific degradation rate between P. chrysogenum ERK1 and Pénicillium frequentans Bi 7/2 (5) indicates that the rates are in the same order of magnitude, between 500-1000 mg phenol/g dry weight/day. The degradation rates obtained for P. chrysogenum ERK 1 are high considering that cultures were carried out under resting mycelium conditions. In addition, the P. chrysogenum strain used in this work was not previously acclimated; for this reason future kinetics studies using the fungus with a previous acclimation period, different Inocula sizes and in the presence of different co-substrates could improve degradation rates.
Leitao et al. (9) described that the degradation ability of P. chrysogenum CLONA2 did not correspond to the visible growth of the mycelia. In this work, the results showed that there is a good correspondence between growth and phenol degradation until biomass remains constant. The same was observed for Aspergillus fumigatus degrading 200 mg/l of phenol (7).
Biomass yield ranges between 2.2932 and 0.2965 (Table 1) depending on the initial phenol concentration. These values are in range with that reported for P. frequentans Bi 7/2 by Hofritcher et al. (9). The results showed that above 200 mg/l, biomass growth remained essentially constant; however P. chrysogenum continues degrading phenol, suggesting that at this stage phenol is used only for energy requirements. Guedes et al. (4) also described that phenol concentration higher than 300 mg/l completely Inhibited fungal growth of P. chrysogenum CLONA 2.
A number of toxic compounds are formed during industrial processes, giving multlcomponent composition of wastewaters. Therefore, the strains used for decontamination processes should not only be highly active to one contaminant but they should also be tolerant of other pollutants or possess different biodégradation abilities (24) and be adaptable to be used in mixed cultures. P. chrysogenum ERK1 showed to produce B-lactam antibiotic. However, when it was grown with phenol as a carbon source It did not show antibiotic production. In addition, the fungus was also able to degrade 2, 4, 6-trichlorophenol (data not shown). For this reason, this fungus could be used in mixed cultures to improve phenol degradation or to contribute to the degradation of multi-substrates.
HPLC analysis detected an Intermediate product with identical retention time to catechol; this result suggests that phenol metabolism could occur via ortho fission of catechol. On the contrary, Leitao et al. (9) could not detect catechol as Intermediary metabolite. However, they showed the presence of hydroquinone but only when the fungus was grown In the presence of phenol and glucose as co-substrate. When the culture had only phenol as carbon source no intermediary metabolites were detected. Marr et al. (12) identified catechol, hydroquinone and cis, cis-muconic acid as intermediary metabolites during the mineralization of phenol by P. simplicissimum SK9117, showing that phenol degradation occurs via the beta-ketoadipate pathway. In the present work, other metabolites were also observed in HPLC chromatograms. However, future studies have to be done to determine their chemical structures.
Catechol dioxigenase activity observed in plate assays agree with HPLC results, suggesting that phenol metabolism could occur via ortho fission of catechol.
Phenolic effluents are being discharged into water bodies and this water can be used for agriculture. These effluents may cause serious environmental problems and health hazards if not treated appropriately. Thus, it is relevant to assess the phytotoxicity of the phenol before and after degradation. To this purpose, the phytotoxicity of plant growing media based on the germination index (Gl) of seeds was evaluated as described by Osma et al. (15). This is one of the most common phytotoxic assays used in the literature.
The Gl combines measurements of relative seed germination and relative root elongation that are both sensitive to the presence of phytotoxic compounds.
Several species have been traditionally used for evaluating phytotoxicity. However there are no standardized seed species in use worldwide (15). For this reason, wheat (Triticum aestivum) was used for this assay because it is a common crop in Argentinean fields. According to Osma et al. (15), Gl values lower than 50 % mean high phytotoxicity, while values between 50 % and 80 % mean moderate phytotoxicity and values over 80 % indicate that the material Is not phytotoxic. Therefore, phenol degradation products were not phytotoxic for wheat.
Finally, the P. chrysogenum strain ERK1 described in the present work degrades phenol as the only source of carbon and energy, with high degradation rates and satisfactory biomass yield under resting mycelium conditions. In addition, phenol degradation products did not show any phytotoxic effects. These characteristics make P. chrysogenum ERK1 attractive to be used in phenol decontamination of soils.
Acknowledgements: this research was supported by Universidad Nacional de Mar del Plata, Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Agencia Nacional de Promoción Científica y Tecnológica (ANPCYT), and MAPFRE foundation. Dr. Wolski, E. A. is a CONICET staff scientist. We are also grateful to Hector L. Villar for providing the wheat seeds.
1. Castellan C, Quadrelli A, Laich F. Surface mycobiota on Argentinean dry fermented sausages. Int J Food Microbiol 2010;142:149-55. [ Links ]
2. Frisvad JC, Samson RA. Polyphasic taxonomy of Pénicillium subgenus pénicillium. A guide to identification of food and air-borne terverticillate penicillia and their mycotoxins. Studies Mycol 2004; 49: 1-174. [ Links ]
3. Glass N, Donaldson G. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 1995; 61: 1323-30. [ Links ]
4. Guedes S, Mendes B, Leitao A. Resorcinol degradation by a Pénicillium chrysogenum strain under osmotic stress: mono and binary substrate matrices with phenol. Biodégradation 2010; 22:409-19. [ Links ]
5. Hofrichter M, Gtinther T, Fritsche W. Metabolism of phenol, chloro- and nitrophenols by the Pénicillium strain Bi 712 isolated from a contaminated soil. Biodégradation 1992; 3:415-21. [ Links ]
6. Huang X, Madan A. CAP3: A DNA sequence assembly program. Genome Res 1993; 9: 868-77.
7. Jones K, Trudgill P, Hopper D. Evidence of two pathways for the metabolism of phenol by Aspergillus fumigatus. Arch Microbiol 1995; 163:176-81. [ Links ]
8. Laich F, Fierro F, Martin JF. Production of penicilli in by fungi growing on food products: identification of a complete penicillin gene cluster in Pénicillium griseofulvum and a truncated cluster in Pénicillium verrucosum. Appl Environ Microbiol 2002; 68:1211-9. [ Links ]
9. Leitao A, Duarte M, Oliveira J. Degradation of phenol by a halotolerant strain Pénicillium chrysogenum. Int Biodeter Biodegrad 2007; 59:220-5. [ Links ]
10. Leitao A. Potential of Pénicillium species in the bioremediation field. Int J Environ Res Public Health 2009; 6:1393-417. [ Links ]
11. Levin L, Papinutti L, Forchiassin F. Evaluation of Argentinean white rot fungi for their ability to produce lignin-modifying enzymes and decolorize industrial dyes. Biores Technol 2004; 94:169-76. [ Links ]
12. Marr J, Kremer S, Sterner O, Anke H. Transformation and mineralization of halophenols by Pénicillium simplicis simum SK9117. Biodégradation 1989; 7:165-71. [ Links ]
13. McErlean C, Marchant R, Banat IM. An evaluation of soil colonisation potential of selected fungi and their production of ligninolytic enzymes for use in soil bioremediation applications. Antonie van Leeuwenhoek 2006; 90:147-58. [ Links ]
14. Nakhla G, Suidan M, Pfeffer J. Control of anaerobic GAC reactors treating inhibitory wastewaters. J Water Pollut Control Fed 1990; 62: 65-72.
15. Osma J, Toca-Herrera J, Rodríguez-Couto S. Transformation pathway of Remazol Brilliant Blue R by immobilised lacease. Biores Technol 2010; 101: 8509-14.
16. Pitt J, Hocking A (editors). Fungi and food spoilage. Maryland, Aspen publisher Inc., 1999.
17. Pitt J (editor). The genus Pénicillium and its teleomorphic states Eupenicillium and Talaromyces. London, Academic Press, 1979.
18. Rodriguez E, Nuero O, Guillén F, Martínez A, Martínez M. Degradation of phenolic and non-phenolic aromatic pollutants by four Pleurotus species: the role of lacease and versatile peroxidase. Soil Biol Biochem 2004; 36:909-16. [ Links ]
19. Rubilar-Araneda O. Biorremediación de suelos contaminados con pentaclorofenol (PCF) por hongos de pudrición blanca. PhD tesis 2007. Universidad de la Frontera, Temuco, Chile.
20. Ruiz-Ordaz N, Ruiz-Lagunez J, Castañón-González J, Hernández-Manzano E, Cristiani-Urbina E, Galíndez-Mayer J. Phenol biodégradation using a repeated batch culture of Candida tropicalis in a multistage bubble column. Rev Latinoam Microbiol 2001 ; 43:19-25.
21. Samson R, Hoekstra E, Frisvad J, Filtenborg O, editors. Introduction to food- and airborne fungi, 6th edition. The Netherlands, Utrecht: Central Bureau Voor Schimmelcultures, 2002.
22. Samson R, Seifert K, Kuijpers A, Houbraken J, Frisvad J. Phylogenetic analysis of Pénicillium subgenus Pénicillium using partial beta-tubulin sequences. Studies Mycol 2004; 49:175-200.
23. Samuels G, Ismaiel A. Trichoderma evansiiand T. Heckfeldtiae. two new T hamatum-Wke species. Mycologia 2009; 101:142-56. [ Links ]
24. Santos V, Linardi V. Biodégradation of phenol by filamentous fungi isolated from industrial effluents: identification and degradation potential. Process Biochem 2004; 39:1001-6.
25. Scow K, Li D, Manilal V, Alexander M. Mineralization of organic compounds at low concentrations by filamentous fungi. Mycol Res 1990; 94: 793-8.
26. Shiftman D, Cohen S. Role of the imp operon of the Streptomyces coelicolor genetic element SLP1 : two imp-encoded proteins interact to autoregulate imp expression and control plasmid maintenance. J Bacterid 1993; 175:6767-74. [ Links ]
27. Tamura K, Dudley J, Nei M, Kumar S. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 2007; 24:1596-9. [ Links ]
28. Yamamoto D, Uchida J. Rapid nuclear staining of Rhizoctonia solani and related fungi with acridine orange and with safranin. Mycologia 1982; 74:145-9. [ Links ]
Recibido: 15/11/2011- Aceptado: 23/4/2011














