Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Revista argentina de microbiología
Print version ISSN 0325-7541
Rev. argent. microbiol. vol.45 no.1 Ciudad Autónoma de Buenos Aires Mar. 2013
MICROBIOLOGÍA BÁSICA
Association between embB mutations and ethambutol resistance in Mycobacterium tuberculosis isolates from Cuba and the Dominican Republic: reproducible patterns and problems
Elba Guerrero1*, Dihadenys Lemus2+, Sergio Yzquierdo2+, Glenda Vílchez1, Mariana Muñoz1, Ernesto Montoro2, Howard Takiff1*
1Instituto Venezolano de Investigaciones Científicas (IVIC), Caracas, Venezuela;
2Instituto de Medicina Tropical Pedro Kourí (IPK), La Habana, Cuba.
*Correspondence. E-mail: htakiff@ivic.gob.ve
ABSTRACT
The relation of ethambutol resistance to embB mutations remains unclear, and there are no reports on ethambutol resistance from the Caribbean. We examined the sequence of embB in 57 distinct Multi-Drug Resistant (MDR) and non-MDR strains of Mycobacterium tuberculosis, mostly from Cuba and the Dominican Republic. embB306 codon mutations were found exclusively in MDR-TB, but in both ethambutol sensitive and resistant strains. Valine substitutions predominated in ethambutol resistant strains, while isoleucine replacements were more common in sensitive strains. Three ethambutol resistant MDR strains without embB306 substitutions had replacements in embB406 or embB497, but these were also found in ethambutol sensitive MDR strains. The results confirm previous findings that amino acid substitutions in EmbB306, EmbB406 and EmbB497 are found only in MDR-TB strains but in both phenotypically resistant and sensitive strains. One ethambutol resistant non-MDR strain did not have any embB mutation suggesting that other undefined mutations can also confer ethambutol resistance.
Key words: Mycobacterium tuberculosis; Ethambutol resistance; Multidrug resistance; embB mutations; Drug sensitivity testing.
RESUMEN
Asociación entre las mutaciones en embB y la resistencia a etambutol en aislamientos de Mycobacterium tuberculosis de Cuba y República Dominicana: patrones y problemas reproducibles. La relación entre resistencia a etambutol y mutaciones en embB no se ha establecido claramente y no existen comunicaciones sobre la resistencia a etambutol en cepas provenientes de países Caribeños. Se evaluó la secuencia del gen embB en 57 cepas de Mycobacterium tuberculosis multi-drogo resistentes (MDR) y no-MDR, la mayoría aislada en Cuba y República Dominicana. Se encontraron mutaciones en el codón embB306 exclusivamente en cepas MDR, tanto en cepas sensibles como resistentes a etambutol. Tres cepas MDR resistentes a etambutol, sin sustituciones en embB306, tenían mutaciones en los codones embB406 o embB497, pero estos cambios se encontraron también en cepas sensibles. Los resultados confirman otros estudios, mostrando que sustituciones aminoacídicas en EmbB306, EmbB406 y EmbB497 se encuentran exclusivamente en cepas MDR, tanto resistentes como sensibles a etambutol. Encontramos una cepa no MDR resistente a etambutol sin ninguna mutación en embB, sugiriendo que otras mutaciones aún no identificadas también pueden conferir resistencia a etambutol.
Palabras clave: Mycobacterium tuberculosis; Resistencia a etambutol; Multi-drogo resistencia; Mutaciones en embB; Prueba de sensibilidad.
+ These authors contributed equally to this publication.
Ethambutol is one of the first line agents used to treat tuberculosis, whose principal target appears to be the arabinosyl transferase EmbB. Although ethambutol has been used for nearly 40 years, there remains uncertainty about the mutations that cause resistance to the drug. The initial studies associating ethambutol resistance with substitutions in EmbB amino acid 306 13 were followed by reports of embB306 mutations in ethambutol susceptible strains 5 and their absence in ethambutol resistant strains 9. Because most ethambutol resistance occurs in multi-drug resistant strains of M. tuberculosis (MDR-TB), it was suggested that embB306 substitutions were not directly associated with ethambutol resistance but rather with MDR-TB 2. However, the causative role of these mutations in ethambutol resistance was subsequently confi rmed when in vitro genetics showed that a susceptible strain can be made resistant to ethambutol by replacing the EmbB 306 methionine with either valine (Met306Val) or isoleucine (Met306Ile), with valine conferring slightly more resistance 14 6, 11. When these mutations are removed from resistant strains by in vitro genetics, the level of resistance decreases.
Adjacent to the embB gene in the M. tuberculosis chromosome are genes encoding two other arabinosyl transferases, embC and embA, which are likely the products of gene duplication 15. Studies of ethambutol resistant strains have described a variety of other mutations in the embB gene, as well as in embC and embA, but the only other substitutions that have been consistently associated with ethambutol resistance are at codons embB406 and embB497 7, 10, 12, 14. Studies with in vitro genetics have demonstrated that substitutions in these amino acids also confer low-level ethambutol resistance 10, but like the embB306 substitution, they have been also found in phenotypically ethambutol-sensitive strains.
We studied a group of 61 ethambutol-resistant and sensitive strains to examine the relationship of embB mutations to ethambutol resistance. The results concisely reproduce the robust fndings in previous studies, and thus serve to summarize the current state of knowledge on mutations associated with ethambutol resistance.
The origins of the 61 clinical strains tested were as follows: 15 were selected from strains isolated from patients in Cuba; 22 were strains selected from patient isolates in the Dominican Republic; 10 strains were sent from the collection of the WHO Supranational Mycobacterial Reference Lab at the Institute for Tropical Medicine (ITM) in Antwerp, Belgium to the PAHO/WHO Collaborating Center in Cuba as part of routine proficiency testing; and 14 strains were isolated in Venezuela. The strains were tested for susceptibility to at least 4 drugs - isoniazid, rifampicin, ethambutol and streptomycin - using the proportion method on Lowenstein-Jensen media (Table 1). The critical concentration for ethambutol resistance with this method was 2 mg/ml, and the results for the ten-strain test panel agreed with the results obtained in the WHO Reference Lab. The fourteen Venezuelan isolates used as negative controls were tested with the proportion method by the Venezuelan National Mycobacterial Reference laboratory and found to be sensitive to the four drugs.
Table 1. Strains used in the study, their sensitivity to four drugs as determined by the proportion method, and the amino acid found at EmbB306
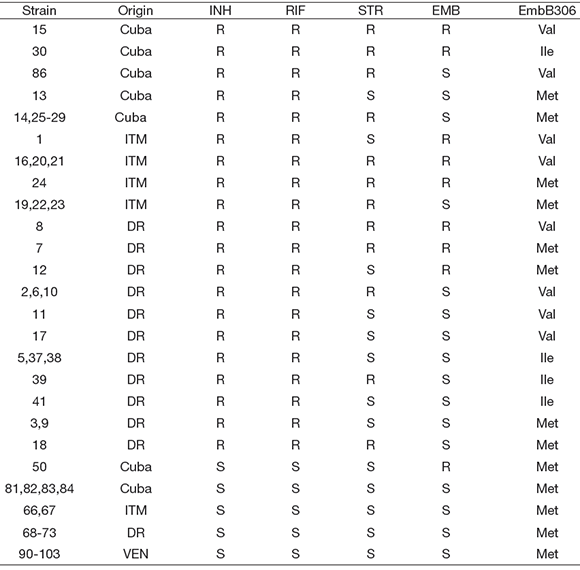
ITM: Institute of Tropical Medicine, Antwerp, Belgium; DR: Dominican Republic; VEN: Venezuela; INH: isoniazid; RIF: rifampicin; STR: streptomycin; EMB: ethambutol.
To eliminate possible bias if more than one strain with the same genotype and phenotype were included in the sample, spoligotyping was performed on all MDR and Venezuelan strains, and those with identical spoligotypes were also analyzed by MIRU15 (1). Isolates with differences in at least two MIRU loci were considered to have different genotypes. Four strains were excluded from the analysis because they were genotypically and phenotypically identical to other strains, so that no strain was represented more than once in the sample. The Venezuelan isolates are representative of that country's most common spoligotypes 1.
Mutations in codon embB306 were detected by amplifying genomic DNA with primers embBF (GGTGCGCGCCATGCCACC) and embBR (GGTCTGGCAGGCGCATCC) to obtain a product of 803 bp 9. To amplify the 908 bp region of embB406-497, we designed primers embBf2 (TTCCTGCTCTG GCATGTCAT) and embBr2 (GGCGTGAACATCAGGAAGAA). To amplify the 1034 bp region upstream of embC, we designed primers embCF1 (CGGCTGGCCCAGGACGTCTA) and embCR1 (GCCGTCGTCGGAGGTGTTGG). The fragments were sequenced with these same primers and also with primer embCR2 (GCACCGGGGTGCTCAGCATT). PCR reactions used 0.02 U/ul Phusion DNA polymerase (Finnzymes), 1X reaction buffer, 5 % DMSO, 0.5 uM each primer, 200 uM each dNTP and 2.5 ul diluted genomic DNA as template. The amplifcation reactions were performed using standard protocols. Annealing temperatures were: 68 °C (embB primers), 59 °C (embC primers) and 56 °C (embBF2/R2 primers). All fragments were sequenced in both directions on an ABI3130XL sequencer in the Genetic and Forensic Studies Unit, IVIC.
Of a total of 30 different MDR strains, only 10 were phenotypically ethambutol-resistant (Table 2), but 53 % (16/30) had EmbB306 substitutions: 70 % (7/10) of ethambutol-resistant MDR and 45 % (9/20) of ethambutol-sensitive MDR strains. Of the ten ethambutol-resistant MDR strains, six had valine substitutions at EmbB306, one had an isoleucine, and three had the wild-type (WT) methionine. All three of the ethambutol-resistant MDR strains without amino acid substitutions at EmbB306 had other embB mutations (see below).
Table 2. Distribution of embB mutations by drug resistance and country of origin

The number and percentage of strains with EmbB306 amino acid substitutions are shown in parentheses on the right. The strains with EmbB406 and EmbB497 substitutions are indicated in brackets. MDR: multidrug- resistant; EMB: ethambutol, ITM: Institute for Tropical Medicine in Antwerp, Belgium.
Of the 20 ethambutol-sensitive MDR strains, 11 were WT at EmbB306, 6 had isoleucine substitutions and 3 had valines. There were no EmbB substitutions in the 27 non-MDR strains, although one was ethambutol-resistant.
Six of the 30 MDR strains had embB mutations at other sites: two had amino acid substitutions at EmbB406 and four at EmbB497 (Table 2). The two Emb406 substitutions were in MDR strains without mutations at embB306: a cysteine substitution for the WT glycine (GGC-TGC) in an ethambutol-resistant strain, and an aspartic acid substitution (GGC-GAC) in an ethambutol-sensitive strain. Of the four strains with mutations at embB497, three had arginine substitutions for the WT glutamine (CAG-CGG) - two in ethambutol-resistant strains with WT Emb306, and one in an ethambutol sensitive strain with WT Emb306. There was one EmbB497 histidine substitution (CAG-CAC) in an ethambutol sensitive strain with isoleucine at EmbB306. The only other mutation found was a synonymous change at codon embB534 (Aspartic acid, GAC-GAT). No mutations were found in the region upstream of embC.
The presence of embB306, 406 and 497 mutations in strains found to be phenotypically sensitive to ethambutol raises doubts about the accuracy of ethambutol sensitivity testing. The level of ethambutol resistance shown by strains constructed in vitro to contain these mutations is generally only 2 - 4-fold higher than that of the parent strains, and ethambutol resistance testing is known to be problematic, yielding MIC values that vary depending upon the culture media and method used 3. However, some clinical isolates have high ethambutol MIC's, well above the low-level of resistance of the in vitro constructed mutants 14, so it is assumed that mutations at additional, unidentified sites, contribute to higher-level resistance 11 and may also be responsible for ethambutol resistance in strains without any mutations in embCAB 9.
Another proposed explanation for the lack of concordance is that some strains with embB mutations are sensitive to ethambutol because these mutations by themselves will not confer phenotypic ethambutol resistance unless accompanied by a second mutation in an as-yet-unknown site 11. The effect of embB substitutions on ethambutol resistance might also vary depending upon other mutations or polymorphisms particular to the genomic background of the strain. Interestingly, eight of the nine ethambutol-sensitive strains with embB mutations were isolated in the Dominican Republic (Table 2). These other putative mutations or strain polymorphisms, could occur in genes coding for proteins upstream in the arabinan biosynthesis pathway, the site of action of the promising new drug benzothiazinone BTZ0434, or in downstream genes encoding proteins that interact with EmbB or are involved in the formation of arabinogalactan.
The results of this study essentially confirm all of the findings consistently reported in previous studies: 1) EmbB306 substitutions are found exclusively in MDR-TB, perhaps due to a proposed advantage these mutations confer for growth in the presence of isoniazid and rifampicin 11; 2) EmbB306 substitutions are present in both ethambutol-sensitive and ethambutol-resistant MDR-strains; 3) valine substitutions at EmbB306 predominate in ethambutol-resistant MDR strains (6/10), while isoleucine substitutions were more common in ethambutol-sensitive MDR isolates (6/20), which is consistent with reports that strains of M. tuberculosis constructed to contain valine in EmbB306 had higher MIC's than strains constructed to contain the isoleucine substitution 11; 4) substitutions at EmbB406 and EmbB497 are also found only in MDR strains and appear to confer ethambutol resistance 10, as they were present in three ethambutol-resistant strains without EmbB306 substitutions, but are also found in ethambutol-sensitive MDR strains; 5) there must be other sites for mutations conferring ethambutol resistance, as some ethambutol-resistant strains do not have any embB mutations. The study also demonstrated that the embB mutations reported in other studies (EmbB306, EmbB406 and EmbB497) are also found in MDR strains from Cuba and the Dominican Republic, a region not represented in prior reports on ethambutol resistance (Table 2).
The detection of embB306/406/497 codon mutations could be a rapid screen for resistance to ethambutol. However, while the sensitivity might be fairly high, it would be hard to predict the specificity of this approach for lack of data on the clinical significance of embB mutations in phenotypically ethambutol-sensitive strains. The only valid criterion for a strain being sensitive or resistant is whether ethambutol can contribute to curing the patient carrying the strain, but this is difficult to assess because ethambutol is always given in combination with several other agents. It could be worthwhile determining the absolute ethambutol MICs for strains that are found to be resistant by screening methods using only a cut-off concentration. Higher doses of ethambutol might be effective against strains with low-level resistance, but might also increase the risk of toxicity 8.
The prospects for developing an accurate and useful molecular screen will depend upon a more complete elucidation of the molecular determinants of ethambutol resistance and the clinical significance of embB mutations in strains that are phenotypically sensitive to ethambutol. Until such information is available, clinicians might consider adding an additional drug or increasing the ethambutol dose to treat patients with "ethambutol-sensitive" MDR isolates carrying EmbB306, 406 or 497 substitutions.
Acknowledgements: the authors thank Dr. Francoise Portaels of the WHO Mycobacterial Reference Lab at the ITM, Antwerp, for providing strains for proficiency testing included in this study, and also Lic. Ana Frías and Lic. Leonarda Reyes of the Laboratorio Nacional de Referencia de Enfermedades Respiratorias (LARNER), Santo Domingo, Dominican Republic, and Dr. Mercedes España and Lic. Carmen Ramírez of the Venezuelan National Tuberculosis Program for providing strains from their respective countries.
Funding: this work was funded by FONACIT (Fondo Nacional de Ciencia, Tecnología y Innovación) Projects G-2005000393 and 20010001851, and by Helmerich & Paine de Venezuela, C.A. through LOCTI (Ley Orgánica de Ciencia, Tecnología y Innovación) project "Las Cepas de Tuberculosis Mas Virulentas de Venezuela," Cuba-Venezuela Project #689, LOCTI project "Tuberculosis en Venezuela: diagnóstico, epidemiología, genética y resistencia a drogas," and CYTED grant 207RT0311.
Competing interests: None declared. Ethical approval: Not required.
1. Abadia E, Sequera M, Ortega D, Mendez MV, Escalona A, Da Mata O, Izarra E, Rojas Y, Jaspe R, Motiwala AS, Alland D, de Waard J, Takiff HE. Mycobacterium tuberculosis ecology in Venezuela: epidemiologic correlates of common spoligotypes and a large clonal cluster defined by MIRU-VNTR-24. BMC Infect Dis 2009; 9: 122. [ Links ]
2. Hazbon MH, Bobadilla del Valle M, Guerrero MI, Varma-Basil M, Filliol I, Cavatore M, Colangeli R, Safi H, Billman-Jacobe H, Lavender C, Fyfe J, Garcia-Garcia L, Davidow A, Brimacombe M, Leon CI, Porras T, Bose M, Chaves F, Eisenach KD, Sifuentes-Osornio J, Ponce de Leon A, Cave MD, Alland D. Role of embB codon 306 mutations in Mycobacterium tuberculosis revisited: a novel association with broad drug resistance and IS6110 clustering rather than ethambutol resistance. Antimicrob Agents Chemother 2005; 49: 3794-802. [ Links ]
3. Madison B, Robinson-Dunn B, George I, Gross W, Lipman H, Metchock B, Sloutsky A, Washabaugh G, Mazurek G, Ridderhof J. Multicenter evaluation of ethambutol susceptibility testing of Mycobacterium tuberculosis by agar proportion and radiometric methods. J Clin Microbiol 2002; 40: 3976-9. [ Links ]
4. Makarov V, Manina G, Mikusova K, Mollmann U, Ryabova O, Saint-Joanis B, Dhar N, Pasca MR, Buroni S, Lucarelli AP, Milano A, De Rossi E, Belanova M, Bobovska A, Dianiskova P, Kordulakova J, Sala C, Fullam E, Schneider P, McKinney JD, Brodin P, Christophe T, Waddell S, Butcher P, Albrethsen J, Rosenkrands I, Brosch R, Nandi V, Bharath S, Gaonkar S, Shandil RK, Balasubramanian V, Balganesh T, Tyagi S, Grosset J, Riccardi G, Cole ST. Benzothiazinones kill Mycobacterium tuberculosis by blocking arabinan synthesis. Science 2009; 324: 801-4. [ Links ]
5. Mokrousov I, Narvskaya O, Limeschenko E, Otten T, Vyshnevskiy B. Detection of ethambutol-resistant Mycobacterium tuberculosis strains by multiplex allele-specific PCR assay targeting embB306 mutations. J Clin Microbiol 2002; 40: 1617-20. [ Links ]
6. Perdigao J, Macedo R, Ribeiro A, Brum L, Portugal I. Genetic characterisation of the ethambutol resistance-determining region in Mycobacterium tuberculosis: prevalence and significance of embB306 mutations. Int J Antimicrob Agents 2009; 33: 334-8. [ Links ]
7. Plinke C, Cox HS, Zarkua N, Karimovich HA, Braker K, Diel R, Rusch-Gerdes S, Feuerriegel S, Niemann S. embCAB sequence variation among ethambutol-resistant Mycobacterium tuberculosis isolates without embB306 mutation. J Antimicrob Chemother 2010; 65: 1359-67. [ Links ]
8. Plinke C, Walter K, Aly S, Ehlers S, Niemann S. Mycobacterium tuberculosis embB codon 306 mutations confer moderately increased resistance to ethambutol in vitro and in vivo. Antimicrob Agents and Chemother 2011; 55: 2891-6. [ Links ]
9. Ramaswamy SV, Amin AG, Goksel S, Stager CE, Dou SJ, El Sahly H, Moghazeh SL, Kreiswirth BN, Musser JM. Molecular genetic analysis of nucleotide polymorphisms associated with ethambutol resistance in human isolates of Mycobacterium tuberculosis. Antimicrob Agents Chemother 2000; 44: 326-36. [ Links ]
10. Safi H, Fleischmann RD, Peterson SN, Jones MB, Jarrahi B, Alland D. Allelic exchange and mutant selection demonstrate that common clinical embCAB gene mutations only modestly increase resistance to ethambutol in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2010; 54: 103-8. [ Links ]
11. Safi H, Sayers B, Hazbon MH, Alland D. Transfer of embB codon 306 mutations into clinical Mycobacterium tuberculosis strains alters susceptibility to ethambutol, isoniazid, and rifampin. Antimicrob Agents Chemother 2008; 52: 2027-34. [ Links ]
12. Shi D, Li L, Zhao Y, Jia Q, Li H, Coulter C, Jin Q, Zhu G. Characteristics of embB mutations in multidrug-resistant Mycobacterium tuberculosis isolates in Henan, China. J Antimicrob Chemother 2011; 66: 2240-7. [ Links ]
13. Sreevatsan S, Stockbauer KE, Pan X, Kreiswirth BN, Moghazeh SL, Jacobs WR Jr., Telenti A, Musser JM. Ethambutol resistance in Mycobacterium tuberculosis: critical role of embB mutations. Antimicrob Agents Chemother 1997; 41: 1677-81. [ Links ]
14. Starks AM, Gumusboga A, Plikaytis BB, Shinnick TM, Posey JE. Mutations at embB codon 306 are an important molecular indicator of ethambutol resistance in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2009; 53: 1061-6. [ Links ]
15. Telenti A, Philipp WJ, Sreevatsan S, Bernasconi C, Stockbauer KE, Wieles B, Musser JM, Jacobs WR Jr. The emb operon, a gene cluster of Mycobacterium tuberculosis involved in resistance to ethambutol. Nat Med 1997; 3: 567-70. [ Links ]
Recibido: 5/6/2012
Aceptado: 29/11/2012













