Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO  uBio
uBio
Compartir
Mastozoología neotropical
versión impresa ISSN 0327-9383versión On-line ISSN 1666-0536
Mastozool. neotrop. vol.22 no.1 Mendoza jun. 2015
NOTA
A new tribe of sigmodontinae rodents (cricetidae)
Ulyses F. J. Pardiñas1, Pablo Teta1*, and Jorge Salazar-Bravo2
1 Instituto de Diversidad y Evolución Austral (IDEAus-CONICET), CC 128, 9120 Puerto Madryn, Chubut, Argentina. [Correspondencia: Ulyses F. J. Pardiñas <ulyses@cenpat-conicet.gob.ar>]
2 Department of Biological Sciences, Texas Tech University, Lubbock, TX-79409, USA.
* Present address: División de Mastozoología, Museo Argentino de Ciencias Naturales “Bernardino Rivadavia,” Avenida Ángel Gallardo 470, C1405DJR Buenos Aires, Argentina.
Recibido 26 noviembre 2014.
Aceptado 25 marzo 2015.
Editor asociado: J Pereira
ABSTRACT.
Phylogenetic hypotheses based on molecular markers indicate that the so-called Reithrodon group, including the extant genera Euneomys, Neotomys, and Reithrodon, formerly within the tribe Phyllotini, is not monophyletic. In turn, a new clade of tribal rank is recovered constituted by Euneomys, Irenomys, and Neotomys, comprising a small, mostly Andean sigmodontine radiation. Within the new clade, here named and diagnosed, Irenomys appears as a taxon with many specialized traits, which suggests an early divergence associated with the exploitation of Nothofagus forests. The living members of the new tribe are characterized by several morphological features including grooved upper incisors, narrow and parallel-sided interorbital regions, enlarged interparietals, and simplified and hypsodont molars. This new phylogenetic scenario implies a high degree of craniodental convergence among several lineages of sigmodontine rodents. The tribal position of several high-crowned extinct forms of sigmodontine rodents (e.g., Panchomys, Tafimys), formerly allocated to the Reithrodon group, is in need of revision.
RESUMEN.
Una nueva tribu de roedores Sigmodontinae (Cricetidae).
Hipótesis filogenéticas basadas en marcadores moleculares indican que el denominado grupo Reithrodon, incluyendo los géneros vivientes Euneomys, Neotomys y Reithrodon, anteriormente dentro de la tribu Phyllotini, no es monofilético. En cambio, un nuevo clado de rango tribal es recuperado, constituido por Euneomys, Irenomys y Neotomys, comprendiendo una pequeña radiación de sigmodontinos mayormente andinos. Dentro de este nuevo clado, aquí nominado y diagnosticado, Irenomys emerge como un taxón con varios rasgos especializados que sugieren una temprana divergencia en el marco de la explotación del bosque de Nothofagus. Los miembros vivientes de la nueva tribu están caracterizados por varios rasgos morfológicos, incluyendo incisivos superiores surcados, regiones interorbitarias angostas y de bordes paralelos, interparietales agrandados y molares simplificados e hipsodontes. Este nuevo escenario filogenético implica un alto grado de convergencia en la morfología cráneo-dentaria entre varios linajes de roedores sigmodontinos. La posición tribal de varias formas extintas de roedores sigmodontinos de coronas altas (e.g., Panchomys, Tafimys), anteriormente ubicadas en el grupo Reithrodon, necesita ser revisada.
Key words: Andean Region; Euneomys; Irenomys; Neotomys; Reithrodon.
Palabras clave: Euneomys; Irenomys; Neotomys; Región Andina; Reithrodon.
The advent of DNA sequence methodology has changed earlier systematic hypotheses, mostly based on morphological features, of mammalian classification (e.g., Meredith et al., 2011). While changes within the Mammalia are perhaps not as impressive as those in other groups of vertebrates (e.g., amphibians; see Frost et al., 2006), many unexpected linkages among taxa have been corroborated leading to new classificatory hypotheses based on molecular markers.
Within the amazingly diverse Sigmodontinae radiation (Patton et al., 2015), the case of the Reithrodon group, an assemblage of medium-sized, stout-bodied, herbivorous rodents previously grouping the living genera Euneomys, Neotomys, and Reithrodon, emerged as paradigmatic. First recognized by Olds and Anderson (1989) as a clade within the tribe Phyllotini, this group was formally diagnosed and strongly supported by the cladistic morphological analyses conducted by Steppan (1993, 1995; see also Braun, 1993). By this time, several extinct species—including, in order of discovery, Panchomys steppani, Ichthyurodon ameghinoi, and Tafimys powelli—were placed within the Reithrodon group, augmenting its known diversity and past distribution (Pardiñas, 1997; Steppan and Pardiñas, 1998; Ortiz et al., 2000). However, only a few years later, several molecular studies questioned both the monophyly of this clade as well as the inclusion of these genera within the tribe Phyllotini. Successive works employing mitochondrial and/or nuclear markers consistently pointed to Reithrodon as a unique linage within Sigmodontinae (e.g., Engel et al., 1998; Smith and Patton, 1999), while Euneomys was recovered as neither close to Reithrodon, nor Phyllotini (e.g., D'Elía, 2003). As a partial result of this changing scenario, Reithrodontini was expanded to include Euneomys and Neotomys plus Reithrodon (Musser and Carleton, 2005). More refined analyses based on interphotoreceptorretinoid binding protein sequences (hereafter IRBP) retrieved a close relationship between Euneomys and the long-tailed, scansorial mouse Irenomys, endemic to the Nothofagus forests of southwestern South America (D'Elía et al., 2006; see also Fabre et al., 2012). Finally, the remaining member of the original Reithrodon group (sensu Olds and Anderson, 1989), the Andean rat Neotomys was recovered as the sister taxa to the Euneomys-Irenomys clade (Martínez et al., 2012; Salazar-Bravo et al., 2013). The close relationship between the terrestrial, stout bodied Euneomys and Neotomys and the forest specialist Irenomys is now well established as a unique branch of the sigmodontine radiation (e.g., Martínez et al., 2012; Salazar-Bravo et al., 2013; Parada et al., 2013; Ventura et al., 2013; Schenk et al., 2013; Pardiñas et al., 2014; see also D'Elía, 2015). This group, however, lacks a formal designation, an issue that we now resolve with the definition and diagnosis of a new tribe of Sigmodontinae (see below).
To assess the definition and diagnosis of the new tribe presented here we follow the concepts portrayed by Reig (1987). In this context we worked based on the extensive morphological revisions produced for several groups of sigmodontine rodents during the last decades (e.g., Voss, 1988; Steppan, 1993, 1995; Pacheco, 2003; Weksler, 2006; Teta, 2013). Several specimens of Euneomys, Irenomys, and Neotomys (see Appendix I) were examined for selected anatomical traits (largely following the characters lists compiled by Weksler [2006]). The obtained product is a combination of morphological traits that is unique to this assemblage, although not formally tested with a cladistic approach. This combination is sufficient to conform an intentional meaning of the polythetic concept of the new clade (Reig, 1987:364). This method was successfully employed in the definition of several tribes recently proposed (e.g., Baiomyini, see Musser and Carleton, 2005; Abrotrichini, see D'Elía et al., 2007) and has the aggregate value of allowing a coordinate advancement of classification and nomenclature, abjuring the claim that this feat is only possible by the identification of unequivocal synapomorphies (a task surely desirable but still under construction). The anatomical terminology used in this contribution—regarding osteological, dental, and soft anatomy—follows that employed by the sources previously mentioned, plus Reig (1977), Carleton (1980), and Hume (1994).
Phylogenetic analyses were conducted with Maximum Likelihood (ML) and Bayesian Inference (BI) on a matrix of 4 nuclear (BRAC1, IRBP, GHR and RAG1) and 1 mitochondrial (Cytochrome b gene; Cytb, thereafter) marker for members of the Sigmodontinae and outgroups (Appendix II). Because the original datasets (as deposited in GenBank) sampled different species of some genera, our dataset included chimeric sequences for some taxa; the effort was made to include data from closely related (in some instances sister taxa) in our alignments; 4 of the 71 ingroup taxa (Sigmodontinae) were represented by chimeric sequences (see Appendix II). Partitioning schemes and substitution models were selected using the Bayesian information criterion in PartitionFinder (Lanfear et al., 2012). The following schemes and substitution models were implemented in the BI analyses: F81+G for GHR_pos2; GTR+G for BRAC1_pos1, BRAC1_ pos2 and RAG1_pos3; GTR+I for GHR_pos3; GTR+I+G for Cytb_pos1, Cytb_pos2, Cytb_ pos3, IRBP_pos1, IRBP_pos2, RAG1_pos1 and RAG1_pos2; K80+G for BRAC1_pos3 and SYM+I+G for GHR_pos1 and IRBP_pos3. Bayesian analyses (100 million generations) were conducted with MrBayes 3.2.2 (Ronquist et al., 2012) on the CIPRES Science gateway (Miller et al., 2010) and with the concatenated dataset. Convergence of the BI analyses was assessed with Tracer v1.6 (Rambaut et al., 2014) to estimate the effective sample sizes for molecular evolutionary parameters (all reached well above 200), and AWTY (Wilgenbusch et al., 2004) to compare posterior probabilities for independent runs (they were, with a correlation coefficient = 0.997) and to test whether 10 subsamples of the chains sampled trees in proportion to their posterior probabilities with the Slide analysis (clade frequencies were very similar). Maximum Likelihood analyses were run with RAxML (Stamatakis et al., 2008) on CIPRES (Miller et al., 2010) under GTR+Gfor all partitions identified by PartitionFinder (see above); invariable sites parameter was not included in any of the analyses as per RaxML manual; the number of alternative runs on distinct starting trees was set to 25 in order to avoid entrapment in local maxima.
Euneomyini, new tribe
Type genus.— Euneomys Coues, 1874.
Contents.— Euneomys Coues, 1874, Irenomys Thomas, 1919, and Neotomys Thomas, 1894
Definition.— The clade composed of the last common ancestor of Euneomys, Irenomys, and Neotomys and all of its descendants.
Diagnosis.— A tribe of the subfamily Sigmodontinae (sensu Reig, 1980), clade Oryzomyalia (sensu Steppan et al., 2004) grouping medium-sized rodents (head and body length 100-160 mm) characterized by the following combination of characters: narrow and parallel-sided interorbital region; well-developed interparietal; distally spatulated hamular processes directly applied on the auditory bullae dorsal borders; enlarged parapterigoyd plates; well excavated sphenopalatine vacuities; short palates; open lacerate foramina; molar toothrows posteriorly divergent; upper incisors grooved by folded enamel/dentine; labial root on M1/m1 absent; internal carotid canal entirely bounded (or nearly so) by the petrosal and ectotympanic portions of the auditory bulla; hypsodont and simplified laminated molars; planar coronal surfaces; procingulum of m1 typically isolated from the remainder portion of the molar; gall bladder present; large caecum (after Sanborn, 1947; Hershkovitz, 1962; Olds and Anderson, 1989; Steppan, 1995; Voss, 1991; Fig. 1 and 2).

Fig. 1. Views of different morphological traits among representative Euneomyini: A. Groove [marker] on incisors (Euneomys mordax; MVZ 183288). B. Narrow and parallel-sided interorbital region (E. mordax; MVZ 183288). C. Internal carotid canal bounded by the petrosal and ectotympanic portions of the auditory bulla (E. chinchilloides; CNP 3641). D. Molar toothrow posteriorly divergent and short palate (Irenomys tarsalis;UACH 2179). E. Well excavated sphenopalatine vacuities and enlarged parapterigoyd plates (E. mordax; MVZ 183288). F. Enlarged interparietal (I. tarsalis;UACH 2179). G. distally spatulated hamular processes directly applied on the auditory bullae dorsal border (E. mordax; MVZ 183288). Abbreviations: as = alisphenoid strut, cc = carotid canal, et = ectotympanic, f = frontal, lf = lacerate foramen, h = hamular process, i = incisor, ip = interparietal, p = palate, pet = pteriotic or petrosal, pp = parapterygoid plate, stf = stapedial foramen, sv = sphenopalatine vacuity.

Fig. 2. Morphology of upper (above) and lower (below) molar occlusal surfaces in Euneomyini: from left to right, Irenomys tarsalis (UACH 2179), Euneomys chinchilloides (CNP 3641), and Neotomys ebriosus (uncatalogued specimen).
Etymology.— The tribal name is formed by adding to the stem of the name of the type genus (ICZN, Article 29.3), Euneomys, the suffix ini (Article 29.2); thus, the family-group name becomes Euneomy + ini = Euneomyini.
Known distribution.— Euneomyini rodents are distributed along the Andes and peri-Andean ranges, including Puna and adjacent lowlands, from central Perú (ca. 10º S) to southernmost South America, reaching Cape Horn islands (ca. 55º 30' S; Musser and Carleton, 2005). By current understanding, there are no areas of sympatry between the genera of Euneomyini. While the range of Euneomys includes both Patagonian lowlands and Andean habitats from southernmost South America to at least 3500 m, Irenomys is mainly confined to the Nothofagus forest. Although Neotomys is also an Andean form (Sanborn, 1947), there is a gap of about 2º in the central Andes where neither Euneomys nor Neotomys have been recorded (Fig. 3).
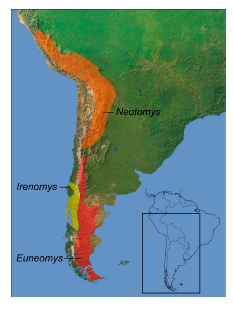
Fig. 3. Generalized distributions of the three genera of Euneomyini.
Biochron.— Late Pleistocene (Euneomys; see Tammone et al., 2014; Neotomys; see Pardiñas and Ortiz, 2001) to Recent. The Pleistocene Euneomys catenatus, a name combination proposed by Hershkovitz (1962) based on Bothriomys catenatus Ameghino, 1889, is a junior synonym of Graomys griseoflavus (cf. Pardiñas, 1995).
Phylogenetic relationships.— The monophyletic group that includes Euneomys, Irenomys, and Neotomys has been recovered with high bootstrap and posterior probability values in all analyses to date. These include the combined analyses of single mitochondrial (Cytb) and nuclear (IRBP) genes (e.g., Martínez et al., 2012; Salazar-Bravo et al., 2013; Parada et al., 2013; Ventura et al., 2013) as well as the combined analyses of multiple nuclear (Schenk et al., 2013) and multiple nuclear and mitochondrial loci (J. Salazar-Bravo, unpublished data); in each of these analyses the topology of this clade is the same: (Irenomys, (Euneomys, Neotomys)).
There remains substantial disagreement as to which taxon (or group) is most closely related to Euneomyini; analyses that include only Cytb and IRBP sequence data tend to suggest that this tribe forms a group (in some cases, with strong Bayesian but poor bootstrap support) with Chinchillula, followed by a clade that includes Andinomys and Punomys (e.g., Salazar-Bravo et al., 2013; Pardiñas et al., 2014) or as a clade within a polytomy that also includes Chinchillula and Reithrodon (see Carrizo and Catalano, in press). Other analyses recover Juliomys as the sister group to Euneomyini (e.g., Ventura et al., 2013: fig. 1), again, often with poor support. Likewise, analyses of 4 nuclear loci suggest that a clade formed by Andinomys and Punomys is the sister group to Euneomyini, although with very low posterior probability and bootstrap below 50% (Schenk et al., 2013); in this topology Chinchillula is recovered as the sister group to the remainder of Oryzomyalia with strong support.
Our own analyses of molecular markers confirm with strong support the position of Chinchillula as the sister group to Oryzomyalia (Fig. 4). In addition, it recovers a polytomy near the base of the Oryzomyalia, with apparent independent origins of the Euneomyini and Andinomys + Punomys clades. Therefore, it appears that, at least for now, the best course of action is to treat each of these clades as independent from each other. It is important to note that a coordinate treatment of the clade composed by Andinomys and Punomys regarding Euneomyini promotes the recognition of a new tribe to contain these 2 Andean genera (Salazar-Bravo et al., in preparation).


Fig. 4. Results of the Bayesian analysis of the combined matrix for members of the Sigmodontinae and outgroups. Numbers indicate posterior probability (left of the diagonal) and ML bootstrap values (right of the diagonal) of adjacent nodes. Dashes indicate nodes that were not recovered in the Maximum Likelihood topology (= -79444.242902). Asterisks represent chimeric sequences (see Appendix II).
Remarks.— The phylogenetic scenario proposed by us and previous workers (e.g., Martínez et al., 2012; Salazar-Bravo et al., 2013; Parada et al., 2013; Schenk et al., 2013) implies a high degree of morphologic convergence among several lineages of sigmodontine rodents. This is particularly evident taking in mind that associations among several of the genera discussed here, including Euneomys, Irenomys, Neotomys, Reithrodon, and Sigmodon, have been repeatedly advanced or discarded (Sanborn, 1947; Hershkovitz, 1962; Pearson and Patton, 1976; Olds and Anderson, 1989). Molecular-based phylogenies are consistent in identifying that at least 4 major groups of tribal rank are necessary to accommodate these genera (Martínez et al., 2012; Salazar-Bravo et al., 2013). Renewed morphological analyses will be required to place correctly the fossil genera Panchomys, Ichthyurodon, and Tafimys.
Within Euneomyini, the genus Irenomys has many specialized traits that appear related to an adaptive exploitation of the Nothofagus forest biome. Paleontological data indicate that the expansion of Nothofagaceae and bamboo (Chusquea), as well as associated micro- to mesothermal families such as Podocarpaceae and Araucariaceae, in Patagonia date from the Middle Eocene-Early Oligocene (cf. Barreda and Palazzesi, 2007). Southern Nothofagus forest, currently restricted to a narrow band in the Andean piedmont between 35º to 55ºS (Ward, 1965), was in the past well represented across interior Patagonia. In addition, molecular data suggest that the genus Irenomys originated between 6.5 Mya (Parada et al., 2013) and 4 Mya (Schenk et al., 2013), which suggests an early and potentially long association with this particular biome. Phenetically, Irenomys is quite divergent with respect to Euneomys and Neotomys. The general bauplan of this Nothofagus endemic shows some convergence with both oryzomyine and some thomasomyine representatives, in particular with those specialized for arboreal life (for example, cladistic analyses based solely on postcranial characters placed Irenomys in the same clade that the arboreal Rhipidomys and Juliomys [Carrizo and Catalano, in press]). Irenomys is a long-tailed form, a trenchant difference to the short-tailed Euneomys and Neotomys, although the 3 share thick, well-haired tails (Fig. 5). A contrasting pattern is also observed in the pes morphology, from the long, narrow and covered by protruding plantar pads in Irenomys to the powerful broad, densely squamated, and long clawed structure in the remainder Euneomyini. Irenomys is an arboreal specialist (Pearson, 1983; Formoso and Sánchez, 2014), although cursoriality is not uncommon in this rat, as the species is readily trapped on the ground (Pearson, 1983). The combined mechanical advantage provided by a long tail and feet covered with fleshy plantar pads to facilitate movements along Nothofagus branches appear to be unequivocal (Carrizo et al., 2013). Differences between Irenomys and the Euneomys-Neotomys pair are not restricted to external traits: the 3 Euneomyini share a well-developed caecum, but that of Irenomys has external taeniae and more complex, pouched structure, not the simple caecum of Euneomys and Neotomys. Cranially, Irenomys lacks the premaxillo-maxillary suture with an acutely angled bend that Olds and Anderson (1989:64) and Steppan (1995:91) identified as synapomorphic for the Reithrodon group.

Fig. 5. External views of Euneomyini representatives, from top to bottom: Euneomys chinchilloides (photo UFJP), Irenomys tarsalis (photo D. Podestá), and Neotomys ebriosus (photo H. Zeballos).
There are a number of character states shared by these genera (Table 1); for example: Irenomys and Euneomys share narrow and high zygomatic plates, which almost completely lack an upper free border, and similar incisive foramina extending to M1 protocones. The “diamond” dental morphology (an example of “transverse lamination” according to Hershkovitz 1962:94) is unique to Irenomys, but a persistent posterofossette on M3 (Fig. 2) is one dental trait shared between Irenomys and Euneomys. An additional dental trait connecting these genera is a lingually displaced anteromedian flexid, notably evident in Irenomys and Neotomys, on the procingulum of the m1; these structures are typically associated with a persistent anteromedian fossetid in Euneomys (Fig. 2). Persistent interfrontal fontanelles have been noted for Irenomys and Neotomys (Gardner and Anderson, 2001), but not in Euneomys. The 3 Euneomyini also share a unilocular-hemiglandular stomach, but this is a widespread condition among sigmodontine rodents (Carleton, 1973). In Neotomys, the cornified portion of the stomach is mostly restricted to the fornix ventricularis while in Irenomys and Euneomys there is more equal distribution between glandular and cornified squamous epithelial portions of the stomach (cf. Carleton, 1973:fig. 3).
Table 1 Morphological character variation among 3 Euneomyini genera.

With the erection of the tribe Euneomyini, incisor grooves acquire a new significance in sigmodontine systematics. The interpretation of this easily diagnostic character was traditionally hampered by poorly resolved phylogenies. Micro-CT scans of upper incisors in Euneomys, Irenomys and Neotomys (see Ohazama et al., 2010:fig. 3) revealed that grooves are caused by folded enamel/ dentine; the same is true for the remainder sigmodontine taxa examined, including living (e.g., Reithrodon, Auliscomys sublimis) and fossil (e.g., Tafimys) representatives (Fig. 6). Although incisor grooves are present in 1 or a few genera in 3 other sigmodontine tribes (Sigmodontini, Reithrodontini, and Phyllotini), their presence characterizes all 3 members of the Euneomyini. Thus, the retention of this trait appears as a signature of the clade and invites further exploration of a potential adaptive role.
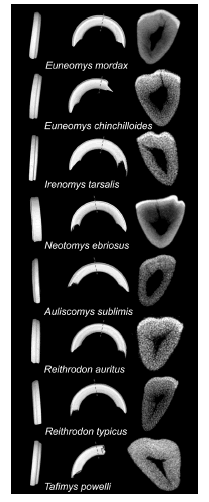
Fig. 6. Folding enamel/dentin in sigmodontine rodent incisors: 3-dimensional reconstructions (frontal and lateral views) and cross-section based on micro-CT scans.
We are at a turning point on the study of the systematics of the Sigmodontinae, as more taxa are incorporated in molecular phylogenetic analyses, although it is also true that key taxa are only recently being included as the result of continuous field work (e.g., Pardiñas et al., 2014). We can expect, therefore, that additional studies will further resolve large parts of the tree, especially with respect to poorly known groups. Nonetheless, we predict that the resolution of the polytomy at the base of the Oryzomyalia will require sequences from many additional loci, perhaps hundreds. This is because gene tree discordance is the expected null when many short branches are combined with large ancestral population sizes (Edwards et al., 2007) such as those that very likely characterized the evolutionary history of the Sigmodontine radiation.
Acknowledgments.
Several of the ideas contained in this note were discussed along many years with G. D'Elía, C. Galliari and P. Ortiz. Micro-CT were kindly and freely processed by J. Blackburn (Dental Institute, Kings College London). D. Podestá and H. Zeballos contributed with photographs of live animals. J. Patton (MVZ) allowed long-term loans of specimens from their collection; D. Flores and M. Gallardo granted the access to MACN and UACH material, respectively. J. Patton acting as reviewer genuinely improved the quality of this manuscript; an anonymous referee also provided some helpful comments. Economic support from Agencia PICT 2008-547 (to UFJP) and the American Philosophical Society and the Systematics Research Fund of the Systematics Association (to JSB) are acknowledged. To the people and institutions herein mentioned goes our deep gratitude. This is the Grupo de Estudio de Mamíferos Australes (GEMA) contribution #14.
LITERATURE CITED
1. BARREDA V and L PALAZZESI. 2007. Patagonian vegetation turnovers during the Paleogene-Early Neogene: Origin of arid-adapted floras. The Botanical Review 73:31-50. [ Links ]
2. BRAUN JK. 1993. Systematic relationships of the tribe Phyllotini (Muridae: Sigmodontinae) of South America. Oklahoma Museum of Natural History, Special Publication, 50 pp. [ Links ]
3. CARLETON MD. 1973. A survey of gross stomach morphology in New World Cricetinae (Rodentia, Muroidea), with comments on functional interpretations. Miscellaneous Publications, Museum of Zoology, University of Michigan 146:1-43. [ Links ]
4. CARRIZO LV and SA CATALANO. In press. First phylogenetic analysis of the tribe Phyllotini (Rodentia: Sigmodontinae) combining morphological and molecular data. Cladistics; DOI: 10.1111/cla.12109. [ Links ]
5. CARRIZO LV, M TULLI, D DOS SANTOS and V ABDALA. 2013. Interplay between postcranial morphology and locomotor types in Neotropical sigmodontine rodents. Journal of Anatomy doi:10.1111/joa.12152. [ Links ]
6. D'ELÍA G. 2003. Phylogenetics of Sigmodontinae (Rodentia, Muroidea, Cricetidae), with special reference to the akodont group, and with additional comments on historical biogeography. Cladistics 19:307-323. [ Links ]
7. D'ELÍA G. 2015. Sigmodontinae incertae sedis. Pp. 70-73 in Mammals of South America, Volume 2 – Rodents (JL Patton, UFJ Pardiñas and G D'Elía, eds.). The University of Chicago Press, Chicago, Illinois.
8. D'ELÍA G, L LUNA, EM GONZÁLEZ, and BD PATTERSON. 2006. On the Sigmodontinae radiation (Rodentia, Cricetidae): An appraisal of the phylogenetic position of Rhagomys. Molecular Phylogenetics and Evolution 38:558-564. [ Links ]
9. D'ELÍA G, UFJ PARDIÑAS, P TETA, and JL PATTON. 2007. Definition and diagnosis of a new tribe of sigmodontine rodents (Cricetidae: Sigmodontinae), and a revised classification of the subfamily. Gayana 71:187-194. [ Links ]
10. EDWARDS S, L LIU, and D PEARL. 2007. High-resolution species trees without concatenation. Proceedings of the National Academy of Science USA. 104:5936–5941.
11. ENGEL SR, KM HOGAN, JF TAYLOR, and SK DAVIS. 1998. Molecular systematics and paleobiogeography of the South American sigmodontine rodents. Molecular Biology and Evolution 15:35-49. [ Links ]
12. FABRE PH, L HAUTIER, D DIMITROV, and E DOUZERY. 2012. A glimpse on the pattern of rodent diversification: A phylogenetic approach. BMC Evolutionary Biology 12:88 doi:10.1186/1471-2148-12-88. [ Links ]
13. FORMOSO A and J SÁNCHEZ. 2014. First description of the breeding nest of Irenomys tarsalis, a sigmodontine rodent endemic to southern Andean forests. Revista Mexicana de Biodiversidad 85:987-989. [ Links ]
14. FROST DR et al. 2006. The amphibian tree of life. Bulletin of the American Museum of Natural History 297:1-370. [ Links ]
15. GARDNER SL and S ANDERSON. 2001. Persistent fontanelles in rodent skulls. American Museum Novitates 3327:1-15. [ Links ]
16. HERSHKOVITZ P. 1962. Evolution of Neotropical cricetine rodents (Muridae) with special reference to the phyllotine group. Fieldiana, Zoology 46:1-524. [ Links ]
17. HUME ID. 1994. Gut morphology, body size and digestive performance in rodents. Pp. 315-323, in: The digestive system in mammals: Food, form and function (DJ Chivers and P Langer, eds). Cambridge University Press, New York. [ Links ]
18. ICZN. 1999. International Code of Zoological Nomenclature, Fourth edition. London: International Trust for Zoological Nomenclature, 117 p. [ Links ]
19. LANFEAR R, B CALCOTT, SY HO, and S GUINDON. 2012. PartitionFinder: Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Molecular Biology and Evolution 29:1695-1701 [ Links ]
20. MARTÍNEZ JJ, LI FERRO, MI MOLLERACH, and RM BARQUEZ. 2012. The phylogenetic relationships of the Andean swamp rat genus Neotomys (Rodentia, Cricetidae, Sigmodontinae) based on mitochondrial and nuclear markers. Acta Theriologica doi:10.1007/ s13364-011-0070-9. [ Links ]
21. MEREDITH RW et al. 2011. Impacts of the Cretaceous terrestrial revolution and KPg extinction on nammal diversification. Science 334 (6055):521-524. [ Links ]
22. MILLER MA, W PFEIFER, and T SCHWARTZ. 2010. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Pp. 1-8, in: Proceedings of the Gateway Computing Environments Workshop (GCE). New Orleans, LA. [ Links ]
23. MUSSER GM and MD CARLETON. 2005. Superfamily Muroidea. Pp. 894-1531, in: Mammal species of the world: A taxonomic and geographic reference (DE Wilson and DM Reeder, eds.). 3rd ed, Johns Hopkins University Press, Baltimore, Maryland. [ Links ]
24. OHAZAMA A et al. 2010. A role for suppressed incisor cuspal morphogenesis in the evolution of mammalian heterodont dentition. Proceedings of the National Academy of Sciences 107:92-97. [ Links ]
25. OJEDA AA, A NOVILLO, RJ OVEJERO, PA CUELLO, C LANZONE, AV BORISENKO, JK BRAUN, MA MARES and RA OJEDA. 2014. A cytogenetic, molecular genetic and morphological study of Patagonian chinchilla mice Euneomys (Rodentia, Cricetidae) in the Southern Central Andes. Mammal Research doi:10.1007/s13364-014-0200-2. [ Links ]
26. OJEDA AA, CA RÍOS, and MH GALLARDO. 2004. Chromosomal characterization of Irenomys tarsalis (Rodentia, Cricetidae, Sigmodontinae). Mastozoología Neotropical 11:95-98. [ Links ]
27. OLDS N and S ANDERSON. 1989. A diagnosis of the tribe Phyllotini (Rodentia, Muridae). Pp. 55-73, in: Advances in Neotropical Mammalogy (KH Redford and JF Eisenberg, eds.). Sandhill Crane Press, Inc., Gainesville, Florida. [ Links ]
28. ORTIZ PE, UFJ PARDIÑAS, and SJ STEPPAN. 2000. A new fossil phyllotine (Rodentia: Muridae) from Northwestern Argentina and relationships of the Reithrodon group. Journal of Mammalogy 81:37-51. [ Links ]
29. PACHECO V. 2003. Phylogenetic analyses of the Thomasomyini (Muroidea: Sigmodontinae) based on morphological data. Doctoral thesis, The City University of New York, New York, USA. [ Links ]
30. PARDIÑAS UFJ. 1995. Sobre las vicisitudes de los géneros Bothriomys Ameghino, 1889, Euneomys Coues, 1874 y Graomys Thomas, 1916 (Mammalia, Rodentia, Cricetidae). Ameghiniana 32:173-180. [ Links ]
31. PARDIÑAS UFJ. 1997. Un nuevo sigmodontino (Mammalia: Rodentia) del Plioceno de Argentina y consideraciones sobre el registro fósil de los Phyllotini. Revista Chilena de Historia Natural 70:543-555. [ Links ]
32. PARDIÑAS UFJ and PE ORTIZ. 2001. Neotomys ebriosus, an enigmatic South American rodent (Muridae, Sigmodontinae): Its fossil record and present distribution in Argentina. Mammalia 65:244-250. [ Links ]
33. PARDIÑAS UFJ, G D'ELÍA, G LESSA, M PASSAMANI, and P TETA. 2014. Nuevos datos morfológicos y una hipótesis filogenética para Phaenomys (Rodentia, Cricetidae). Mastozoología Neotropical 21:251-261. [ Links ]
34. PARADA A, UFJ PARDIÑAS, J SALAZAR-BRAVO, G D'ELÍA, and E PALMA. 2013. Dating an impressive Neotropical radiation: Molecular time estimates for the Sigmodontinae (Rodentia) provide insights into its historical biogeography. Molecular Phylogenetics and Evolution 66:960-968. [ Links ]
35. PATTON JL, UFJ PARDIÑAS, and G D'ELÍA. 2015. Mammals of South America, Volume 2 - Rodents. The University of Chicago Press, Chicago, Illinois. [ Links ]
36. PEARSON OP. 1983. Characteristics of a mammalian fauna from forests in Patagonia, southern Argentina. Journal of Mammalogy 64:476-492. [ Links ]
37. PEARSON OP and JL PATTON. 1976. Relationships among South American phyllotine rodents based on chromosome analysis. Journal of Mammalogy 57:339-350. [ Links ]
38. REIG OA. 1980. A new fossil genus of South American cricetid rodents allied to Wiedomys, with an assessment of the Sigmodontinae. Journal of Zoology (London) 192:257-281. [ Links ]
39. REIG OA. 1987. An assessment of the systematics and evolution of the Akodontini, with the description of new fossil species of Akodon (Cricetidae: Sigmodontinae). Pp. 347-399 in Studies in Neotropical mammalogy – Essays in honor of Philip Hershkovitz(BD Patterson and RM Timm, eds.). Fieldiana, Zoology, New Series 39.
40. RONQUIST F, M TESLENKO, P VAN DER MARK, DL AYRES, A DARLING, S HOHNA, B LARGET, L LIU, MA SUCHARD, and JP HUELSENBECK. 2012. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology 61:539-542. [ Links ]
41. RAMBAUT A, MA SUCHARD, D XIE, and AJ DRUMMOND. 2014. Tracer v1.6, Available from http://beast.bio.ed.ac.uk/Tracer [ Links ]
42. SALAZAR-BRAVO J, UFJ PARDIÑAS, and G D'ELÍA. 2013. A phylogenetic appraisal of Sigmodontinae (Rodentia, Cricetidae) with emphasis on phyllotine genera: Systematics and biogeography. Zoologica Scripta 42:250-261. [ Links ]
43. SANBORN CC. 1947. The South American rodents of the genus Neotomys. Fieldiana, Zoology 31:51-57. [ Links ]
44. SCHENK JJ, KC ROWE, and SJ STEPPAN. 2013. Ecological opportunity and incumbency in the diversification of repeated continental colonizations by muroid rodents. Systematic Biology 62:837-864. [ Links ]
45. SMITH MF and JL PATTON. 1999. Phylogenetic relationships and the radiation of sigmodontine rodents in South America: Evidence from cytochrome b. Journal of Mammalian Evolution 6:89-128. [ Links ]
46. STEPPAN SJ. 1993. Phylogenetic relationships among the Phyllotini (Rodentia: Sigmodontinae) using morphological characters. Journal of Mammalian Evolution 1:187–213.
47. STEPPAN SJ. 1995. Revision of the tribe Phyllotini (Rodentia: Sigmodontinae), with a phylogenetic hypothesis for the Sigmodontinae. Fieldiana, Zoology, New Series 80:1-112. [ Links ]
48. STEPPAN SJ and UFJ PARDIÑAS. 1998. Two new fossil muroids (Sigmodontinae: Phyllotini) from the Early Pleistocene of Argentina: Phylogeny and paleoecology. Journal of Vertebrate Paleontology 18:640-649. [ Links ]
49. STEPPAN SJ, RM ADKINS, and J ANDERSON. 2004. Phylogeny and divergence-date estimates of rapid radiations in muroid rodents based on multiple nuclear genes. Systematic Biology 53:533-553. [ Links ]
50. TAMMONE M, A HAJDUK, P ARIAS, P TETA, E LACEY, and UFJ PARDIÑAS. 2014. Last Glacial Maximum environments in northwestern Patagonia revealed by fossil small mammals. Quaternary Research 82:198-208. [ Links ]
51. TETA P. 2013. Relaciones filogenéticas de la tribu Abrotrichini (Rodentia, Cricetidae): análisis separados y combinados de evidencias morfológicas y moleculares. Doctoral thesis, Facultad de Ciencias Naturales y Museo, Universidad Nacional de La Plata, La Plata, Buenos Aires, Argentina. [ Links ]
52. STAMATAKIS A, P HOOVER, and J ROUGEMONT. 2008. A rapid bootstrap algorithm for the RAxML Web Servers. SystematicBiology57:758-771. [ Links ]
53. VENTURA K, MJ SILVA, L GEISE, Y LEITE, UFJ PARDIÑAS, Y YONENAGA-YASSUDA, and G D'ELÍA. 2013. The phylogenetic position of the enigmatic Atlantic forest-endemic spiny mouse Abrawayaomys (Rodentia: Sigmodontinae). Zoological Studies 52:55 doi:10.1186/1810-522X-52-55. [ Links ]
54. VOSS RS. 1988. Systematics and ecology of Ichthyomyinae rodents (Muroidea): Patterns of morphological evolution in a small adaptative radiation. Bulletin of the American Museum of Natural History 188:259-493. [ Links ]
55. VOSS RS. 1991. An introduction to the Neotropical muroid rodent genus Zygodontomys. Bulletin of the American Museum of Natural History 210:1–113.
56. WARD RT. 1965. Beech (Nothofagus) forests in the Andes of Southwestern Argentina. The American Midland Naturalist 74:50-56. [ Links ]
57. WEKSLER M. 2006. Phylogenetic relationships of oryzomine rodents (Muroidea: Sigmodontinae): separate and combined analyses of morphological and molecular data. Bulletin of the American Museum of Natural History 296:1-149. [ Links ]
58. WILGENBUSCH JC, DL WARREN, and DL SWOFFORD 2004. AWTY: A system for graphical exploration of MCMC convergence in Bayesian phylogenetic inference. http://ceb.csit.fsu.edu/awty. [ Links ]
Specimens examined, including dried specimens as well as fluid preserved animals, are housed in the following mammal collections: Colección de Mamíferos del Centro Nacional Patagónico (CNP; Puerto Madryn, Chubut, Argentina), Colección de Material de Egagrópilas y Afines “Elio Massoia” (CNP-E; Puerto Madryn, Chubut, Argentina); Colección Nacional de Mastozoología, Museo Argentino de Ciencias Naturales “Bernardino Rivadavia” (MACN; Buenos Aires, Argentina), Museum of Vertebrate Zoology (MVZ; Berkeley, California, USA).
Euneomys chinchilloides (n = 27). Argentina: Chubut, cercanías de Esquel (CNP 4), Cañadón de la Madera, Sierra de Tepuel (CNP 2388, 2422-2424), Altiplanicie del Somuncura, Laguna Blanca (CNP 2406-2421); Establecimiento La Maroma (CNP 4420, 4429, 4621); Santa Cruz, nacimiento río Ecker (CNP 3674), estancia La Ensenada (CNP 2427, 3641).
Euneomys mordax (n = 2). Argentina: Neuquén, 1.5 km S Copahue (town) (MVZ 183288, 183289).
Irenomys tarsalis (n = 7). Argentina: Depto. Huiliches, Arroyo de las Tierras Coloradas, 1 km E and 2.4 km S Cerro del Chivo (MVZ 163459); Neuquén, Depto. Los Lagos, 3 km NW Confluencia (MVZ 159421), Pilolil (CNP 730) Loma Redonda, Ruca Hué, San Martín de los Andes (CNP 3642); Chubut, lago La Plata (CNP 731); Río Negro, Alto Río Villegas (CNP 778). Chile: Los Lagos, Osorno, Valle de La Picada, 715 m (UACH 2179).
Neotomys ebriosus (n = 35). Argentina: Salta, San Antonio de los Cobres (CNP 3639, 3640); San Juan, San Guillermo (30 uncatalogued from owl pellets). Perú, Puno, 13 mi ENE Crucero, Abra Aricoma (MVZ 139590); Arequipa, 2 km W Sumbay (MVZ 174043).
Taxon sampling and access codes for 4 nuclear (BRAC1, IRBP, GHR and RAG1) and 1 mitochondrial (Cytb) markers used in the phylogenetic analyses conducted in this paper.

















