Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares em
SciELO
Similares em
SciELO  uBio
uBio
Compartilhar
Biocell
versão impressa ISSN 0327-9545
Biocell vol.35 no.1 Mendoza jan./abr. 2011
ORIGINAL ARTICLES
Detection of single copy sequences using BAC-FISH and C-PRINS techniques in sunflower chromosomes
Paola Talia1, Eduardo J. Greizerstein1, H. Esteban Hopp1,2, Norma Paniego1, Lidia Poggio2 and Ruth A. Heinz1,2*
1. Instituto de Biotecnología, CICVyA, INTA Castelar, N. Repetto y Los Reseros s/n, 1686 Hurlingham, Provincia de Buenos Aires, Argentina.
2. Departamento de Ecología, Genética y Evolución and Departamento de Fisiología, Biología Molecular y Celular, Facultad de Ciencias Exactas y Naturales, Universidad de Buenos Aires, Ciudad Universitaria, Pabellón II, C1428EHA, Buenos Aires, Argentina.
*Address correspondence to:
Ruth Heinz.
Instituto de Biotecnología CICVyA / INTA. Dr. N. Repetto y Los Reseros. (1686) Hurlingham, Buenos Aires. Argentina. Fax: (54-11) 4481 2975. Tel.: (54-11) 4621 1447 ó 0199. E-mail: rheinz@cnia.inta.gov.ar
Abstract: Bacterial artificial chromosome - fluorescence in situ hybridization (BAC-FISH) and cycling-primed in situ labeling (C-PRINS) techniques were evaluated for integration of physical and genetic maps of sunflower (Helianthus annuus L.). Single-site SSR markers were selected from three linkage groups of a high-density sunflower genetic map. This selection was based on previously identified QTL associated to S. sclerotiorum. These markers were used to select BACs contaning single copy sequences for BAC-FISH aplication. Blocking of highly dispersed repetitive sunflower sequences reduced unspecific hybridization, and allowed the detection of specific signals for BACs containing SSR markers HA4222 and HA2600, anchored to LG 16 and LG 10, respectively. Single-site FISH signal detection was optimized by adjusting the relative quantity and quality of unlabelled repetitive sequences present in the blocking DNA. The SSR marker ORS1247 anchored to the LG 17 was detected by C-PRINS, which yielded fluorescence signals that were specific and intense. This progress in localizing single-copy sequences using BAC-FISH and indirect C-PRINS strategies in sunflower will facilitate the integration of genetic and physical maps, allowing the identification of chromosomes containing key genes and/or QTL associated to agronomic important traits in sunflower.
Key words: Sunflower; BAC-FISH; C-PRINS; Single-copy sequences; Physical mapping
Introduction
Sunflower (Helianthus annuus L.) is the third most important source of edible vegetable oil worldwide. It belongs to the Asteraceae family which includes over forty economically important crops, such as lettuce, chamomile and chicory among others. It is a diploid species with 2n=2x=34 with an estimated genome size of about 3,500/1C Mb (Baack et al., 2005).
In the last decade, significant progress has been made in sunflower genomics regarding the development of a large number of molecular markers such as SSR or Simple Sequence Repeat (SSR) (Paniego et al., 2002; Tang et al., 2002, 2003), Expressed Sequence Tags (EST) databanks (http://cgpdb.ucdavis.edu/sitemap.html; www.genoplante.com (INRA Member Group, France); (Gentzbittel et al., 1999; Fernández et al., 2003; Heesacker et al., 2008) and functional markers derived from ESTs as SNPs (Lai et al., 2005; Liu and Burke, 2006; Kolkman et al., 2007; Fusari et al., 2008), SSR-ESTs and Indels (Kumpatla and Mukhopadhyay, 2005; Pashley et al., 2006; Heesacker et al., 2008). In addition, the construction of several genetic linkage maps including both neutral and functional markers (Tang et al., 2002, 2003; Yu et al., 2003; Al-Chaarani et al., 2004; Lai et al., 2005; Kiani et al., 2007) as well as the development of several BAC libraries (Gentzbittel et al., 2002; Özdemir et al., 2004; Bouzidi et al., 2006; Feng et al., 2006) have been reported. In spite of these advances, the association between linkage groups (LGs) and chromosomes has not been established yet.
Reliable cytological techniques as fluorescence in situ hybridization (FISH) and primed in situ labelling (PRINS) represent key tools for association of physical and genetic maps. FISH has been the method of choice for mapping repetitive sequences and thus, enabling the characterization of the chromosome complement in different species (Minelli et al., 2000; Ceccarelli et al., 2007). However, the application of these strategies to localize single locus DNA sequences has been hampered by technical difficulties related, at least in part, to the use of rather short sequences as probes (Danilova and Birchler, 2008). The use of probes containing few kilobases of complementary DNA leads to lower sensitivity in conventional FISH assays, particularly in applications involving large genome size plants like sunflower. This technical difficulty can be overcome by using large-insert genomic clones like BACs (Bacterial Artificial Chromosomes) as probes in FISH (Jiang et al., 1995; Lapitan et al., 1997; Desel et al., 2001; Islam-Faridi et al., 2002; Koorneef et al., 2003; Lee et al., 2003) leading to the strategy known as BAC-FISH (Shizuya et al., 1992). However, the identification of individual BAC clones suitable for FISH is not trivial because of the frequent presence of repetitive DNA in these clones that may jeopardize the detection of specific unique sequences. The use of overlapping oligonucleotides (overgo) for identification of BACs containing specific-unique-site-sequences, has been successfully reported for different plant species, including sunflower (Chen et al., 2002; Gardiner et al., 2004; Yüksel and Paterson, 2005; Feng et al., 2006).
PRINS technique (Koch et al., 1989) represent an alternative to the use of FISH for localization of DNA sequences in plant chromosomes and it has also been reported for different plant species (Menke et al., 1998; Kubaláková et al., 2001). This strategy was found to be particularly useful for detection of high-copy tandem repeats. However, for the detection of low-copy repeats, a more sensitive variant of PRINS named cycling PRINS (C-PRINS), which involves serial thermal cycles analogous to polymerase chain reaction (Gosden et al., 1991; Kubaláková et al., 2001), was recommended. Compared to FISH, PRINS has several advantages including the minimal sequence information required for oligonucleotide primer design, the easiest and fastest detection of the technique based on primer hybridization to homologous targets within densely structured chromatin, compared to the rather large probes commonly used for FISH, the avoidance of probe labeling and the possibility to perform the primer extension even into flanking scattered repetitive sequences, thus strengthening the signal for a short unique sequence (Menke et al., 1998).
In the present work we report the use of the BAC-FISH and C-PRINS techniques for the detection of single-copy DNA sequences on sunflower chromosomes in order to evaluate the potential of both techniques for further applications in the integration of physical and genetics maps.
Materials and Methods
Selection and screening of BAC clones for FISH
The BAC clones used in this study were selected from an 8.3x genome coverage sunflower BAC library with an average insert size of 125 kbp (BAC: HA_HBa. www.genome.clemson.edu).
Unique genomic sequence flanking 32 different microsatellites selected from a high-density genetic map of sunflower and anchored to LG 9, 10 and LG 16, (Kiani et al., 2007) were used to design overlapping oligonucleotide probes (http://genome.wustl.edu/gsc/overgo/overgo.html) (McPherson et al., 2001).
Overgos were designed using the Overgo 1.02i program (http://www.mouse-genome.bcm.tmc.edu/webovergo/OvergoDescription.asp) and oligoSpawn software (http://oligospawn.ucr.edu). Overgos were synthesized and then individually labelled with [α-32P]dATP and [α-32P]dCTP, according to the method developed by J. D. McPherson (http://www.tree.caltech.edu). Pairs of overgo primers (10 pmol for each primer) were added to 3.5 μl H2O and denatured at 80°C for 5 min, followed by 10 min at 37°C and afterwards cooled on ice. Heat-denatured overgo primer pairs were mixed with 0.1 mg/µl BSA, 2 µl OLB (oligo labeling buffer without dATP, dCTP, and random hexamers), 2 U Klenow DNA polymerase fragment (Invitrogen, USA), 0.5 µl [α-32P]dATP (3,000 Ci/mmol) and 0.5 µl [α-32P]dCTP (3,000 Ci/mmol), in a final 10 µl reaction volume. Subsequently, the reaction mixture was incubated for 1 h at 37ºC, followed by a 10 min denaturing step performed at room temperature, with the addition of NaOH at 0.2 N final concentration. Removal of unincorporated radioactive nucleotides was performed using "QiAquick® Nucleotide Removal Kit" (Qiagen, Germany).
Two BAC-library high-density filters were screened with pooled 10 to 12 overgo probes.
Filters were prehybridized in sodium polyanethol sulfonate hybridization buffer (PAES) at 60°C for 4 h. Labelled probes were denatured and then added to the hybridization tubes and then allowed to hybridize overnight at 60°C. The filters were washed in a series of buffers with increasing stringency: 2X SSC and 0.1% SDS at 60°C for 5 min, 1.5X SSC and 0.1% SDS at 60°C for 15 min, 1X SSC and 0.1% SDS at 60°C for 15 min, and 0.5X SSC and 0.1% SDS at 60°C for 15 min (Sambrook et al., 1989). Filters were sealed in plastic sheets, exposed for 72 h and then scanned with a Typhoon trio (Amersham Biosciences, UK) using the program ImagenQuant (Amersham Biosciences, UK).
PCR screening, BAC DNA purification and probe labeling
DNAs from BAC positive clones were purified using NucleoBond® Xtra Midi Plus (Machery-Nagel, Germany). PCR verification of positive clones was carried out using SSR specific primers and 80 ng of DNA from each clone identify in the screening in the presence of 0.25 mM of each dNTP, 1.5 mM MgCl2, 0.75 U of Taq DNA polymerase (Invitrogen, USA), 1X Taq polymerase buffer and 0.25 µM of each primer, in a final volume of 25 µl. Following an initial 5 min denaturalization at 95°C for, 35 PCR cycles of 95°C for 40 s, 62°C for 40 s and 72°C for 2 min and a final extension at 72°C for 10 min were performed. PCR products were separated using 2% standard TAE agarose gel electrophoresis.
DNAs from BAC were labelled with digoxigenin-11-dUTP by random priming labeling reaction (Boehringer Mannheim, Germany) and used as probes for fluorescence in situ hybridization (FISH).
Analysis of repetitive/transposon content in BAC clones using repetitive DNA as probes
Three repetitive DNA regions previously described in sunflower were amplified using genomic DNA from the inbred line HA89 and the following specific primers: SSR-Ha785 (GenBank AN: GF100475) with similarity to a highly repetitive sequence family of Helianthus annuus (GenBank AN: AJ009965) was amplified using forward (5' GTCTCGGGATCCAAGATTGA 3') and reverse (5' TCACGACGGTTGTAAAACGA 3') primers (Paniego et al., 2002); HaRetro3 (GenBank AN: DQ229838.1) with similarity to a gypsy-like retrotransposon family, was amplified using forward (5' AGGGCATTCAAATGGCTATG 3') and reverse (5' GTCTCATCCGGAAGATCCAA 3') primers (Tang et al., 2006) and HaRep1 (GenBank AN: AJ009967) with similarity to a copia-like retrotransponson family, was amplified using forward (5' TCTCAGAACCTCGGCAATCT 3') and reverse (5' GGCAGCAAAAGAGAAAATG 3') primers (Santini et al., 2002).
PCR reaction containing 100 ng of HA89 DNA and the following reagents: 10 mM PCR buffer, 0.25 µM of each specific primer, 0.2 mM of dNTPs, 0.75 U of Taq polymerase (Roche, France), was conducted in an Eppendorf thermocycler, with an initial denaturing step of 3 min at 94ºC followed by 35 cycles of 1 min at 94ºC, 1 min at 55ºC, 30 s at 72ºC, and a final extension step of 10 min at 72ºC. PCR amplification products were run at 80 V on 1% agarose gel electrophoresis and visualized by staining with ethidium bromide. The corresponding bands were excised and purified with Qiaex II gel extraction kit (QIAgen, Germany). Probes were labelled with [α-32P]-dCTP by random priming according to the manufacturer's recommendations (Promega Biotech, USA). These highly repetitive DNA fragments were used as probes in Southern blot hybridization assays to characterize the single-locus microsatellite containing BAC clones and to block unspecific hybridization in BAC-FISH experiments (see bellow).
Southern-blot hybridization
Positive clones were isolated and purified using a QIAgen Plasmid Midi Kit (QIAgen, Germany) and digested with EcoRI restriction enzyme, electrophoresed in 0.8% agarose gels and blotted to nylon membranes Hybond N+ (GE Healthcare, UK) as described by Sambrook et al.(1989). The filters were prehybridized using a buffer containing PAES at 65ºC for 16 h. Denatured [α-P32]-labelled probes (either Ha785, HaRep1 or HaRetro3 amplified sequences) were added and allowed to hybridize overnight at 65ºC, during the course of separate assays. Filters were washed in a series of buffers with increasing stringency: 2X SSC and 0.1% SDS at room temperature for 20 min, 1.5X SSC and 0.1% SDS at room temperature for 15 min, and 0.5X SSC and 0.1% SDS at 65ºC for 15 min (Sambrook et al., 1989). Filters were sealed in plastic bags, exposed for three days to X-rays sensitive screens, and then scanned with a Typhoon trio scanner (Amersham Biosciences, UK).
Chromosome preparations
Inbred line HA89 was selected for chromosome preparations. Root tips, obtained from seeds germinating over moisturized paper, were treated with 0.05% colchicine for 2 h at room temperature, fixed in absolute ethanol/acetic 3:1 (v/v) and stored at 4°C for several weeks. For cell wall digestion, root tips were treated with in an enzymatic solution (2% cellulose [w/v] plus 20% pectinase [v/v]) for 3 h at 37°C and stored in 0.01 M citric acid-sodium citrate buffer at 4°C for 24 h. Digested material was transferred to a slide containing a single drop of 45% acetic acid and flamed before squashing.
Fluorescence in situ hybridization (FISH)
FISH was conducted according to the methods described by Schwarzacher and Heslop-Harrison (2000). Chromosome slides were incubated 10 min in 3:1 (v/v) ethanol: acetic acid and then washed twice in absolute ethanol for 10 min.
Chromosome slide preparations were pre-treated with RNase for 1 h at 37°C and fixed with 4% paraformaldehyde. Hybridization mixture, 30 μl per slide, was prepared by adding 50% deionized formamide, 10% dextran sulphate, 2X SSC (0.3 M NaCl, 0.03 M Na3-citrate), 0.1% SDS (sodium dodecyl sulphate), 0.3 mg/ml of sheared salmon sperm DNA (Sigma, USA), 100 ng of probe and 10 to 80 times of blocking DNA. Taking into account the outcome of BAC-Southern blot assays, DNA fragment from SSR-Ha785 and / or HaRep1 were used as blocking DNA. Hybridization mixture was denatured at 70ºC for 15 min. After chilling on ice for 5 min, 30 μl of the denatured mixture was applied to each slide and hybridization was performed using a thermocycler (Eppendorf Mastercycler, Germany) at 75°C for 7 min, 55-60°C 30 s, 45°C 5 min, 38°C 5 min and 37°C 10 min and then incubated overnight at 37°C in a humid chamber. Stringent washes were made in 2X SSC at 42°C for 10 min, 20% formamide/0.1X SSC at 42°C for 10 min, followed by two washes in 0.1X SSC at 42°C for 5 min, 2X SSC at 42°C for 5 min, 4X SSC/0.2% Tween at 42°C for 5 min and 4X SSC/0.2% Tween at room temperature for 5 min. Slides were blocked for 5 min at room temperature with 5% bovine serum albumin (BSA) (w/v) in 4X SSC/0.2% Tween.
Detection of digoxigenin hybridization sites was carried out by incubating the slides in a solution of 4 μg/ml of anti-digoxigenin-FITC fluorescein isothiocyanate (Roche, France) in 5% BSA (w/v) in 4X SSC/0.2% Tween for 1 h at 37°C. Afterwards, slides were washed in 4X SSC/0.2% Tween three times for 10 min each. Immediately, DNA was stained with 4 μg/ml DAPI (4', 6'-diamidino-2-phenylindole) and washed briefly with 4X SSC/Tween 0.2%. After rinsing, Vectashield antifade solution (Vector Laboratories, USA) was applied.
Cycling-Primed in situ labeling (C-PRINS)
PCR oligonucleotide primers from single locus microsatellite ORS1247 (Accession number: BV006602), anchored in the LG 17, of 339 pb in lenght was selected to carry out C-PRINS. This SSR was selected based on a previously identified QTL associated to S. sclerotiorum (Maringolo, personal comunication).
C-PRINS technique was conducted according to methods described by Kubaláková et al. (2001) with few modifications. Extension mix was performed in a volume of 50 μl containing 1X DNA Taq polymerase buffer (Invitrogen, USA), 2.5 mM MgCl2, 2 μM of each primer, 100 μM of dATP, dCTP and dGTP, 34 μM dTTP, 8 μM digoxigenin-11-dUTP (Roche Applied Science, France), 2 U Taq DNA polymerase and deionized H2O.
Glass slides were sealed within a frame chamber (Bio-Rad, USA). Subsequently, 30 μl of reaction mix was added over the delimited area and a polyester coverslip was placed over it. Slides were then placed on a thermocycler (Eppendorf, Mastercycler, Germany) and subjected to the following amplification program: 91°C 4 min followed by 35 cycles of 91°C for 45 s, 55-60°C for 45 s and 72°C for 45 s and, at last, 72°C for 10 min.
Detection of the reaction was carried out by adding 100 μl of stop buffer (2.923 g of NaCl, 1.861 g of Na2EDTA in 100 μl of deionized H2O, pH 8) and incubated 2 min at 70°C. One hundred μl of BSA were added to each preparation, which was later incubated with 50 μl of anti-Dig fluorescein at 37°C for 30 min.
Slides were washed with 4XSSC/0.2% Tween for 5 min at room temperature for three times and later counterstained with 75 μl of DAPI (2 μg/ml) for 15 min. Chromosome preparations were mounted in Vectashield antifade solution (Vector Laboratories, USA).
Image analysis
Slides were examined with an epifluorescence microscope (Leica DMLB, Germany). Individual images obtained from each filter were captured using a Leica DFC 350 FX camera. These images were optimized for best contrast and brightness with Adobe Photoshop CS2 program.
Results
In order to evaluate BAC-FISH and C-PRINS methods, SSR markers targeting three LGs of a high-density sunflower genetic map (Kiani et al., 2007) were chosen. LGs 10, 16 and 17 were selected based on previous identification of QTL for resistance to S. sclerotiorum (Maringolo, 2007).
Different groups of probes, each containing 10-12 overgoes were used for hybridization to BAC filters as described in Materials and Methods. More than 70 positive clones were obtained, out of which, 35 were selected for further analysis. Six BAC clones representing three linkage groups were confirmed by PCR directed to the SSR tags and reconfirmed by means of Southern blot hybridizations (Table 1).
Table 1. Name, type and repetition motif of SSRs contained in the BAC clones used for BAC-FISH in sunflower chromosomes.

When each one of the six BAC clones were used as probe in FISH experiments, hybridization signals were found scattered over all the chromosomes (data not shown). This result strongly suggests that these BACs contain samples of the repetitive DNA sequences scattered within the sunflower chromosomes.
These six clones were characterized using Southern blot by hybridization with different repetitive sequences (Table 1). This evaluation confirmed that the hybridization patterns observed in FISH experiments were due to transposable elements present in each BAC clone. Among these 6 clones, BAC2 and BAC5 showed the highest repetitive element copies particularly when hybridized with SSR-Ha785 probe (Fig. 1a). When the HaRep1 sequence, which showed similarity to copia-like retrotransposons (Santini et al., 2002), was used as probe, strong hybridization signals were detected in BAC1 and BAC6 clones (Fig. 1b). On the other hand, no signals were observed when a probe with similarity to gypsy-like retrotransposon was used (Tang et al., 2006) (data not shown).
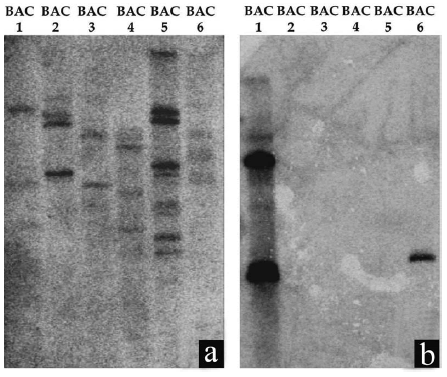
Figure 1. BAC-Southern blot analysis. Positive BAC clones were isolated, purified, EcoRI digested, electrophoresed, blotted to nylon membranes and hybridized to [a-P32]-labelled probes. (a) Hybridization signals observed in the six BACs clones using SSR-Ha785 as probe. (b) Hybridization signals observed in BAC 1 and BAC 6 clones using HaRep1 as probe.
Based on these results, two BAC clones (BAC3 and BAC6) were selected to perform BAC-FISH using the repetitive sequences SSR-HA785 and / or the HARep1 as blocking DNA.
Specific signal were obtained in both cases and confirmed after thorough observation of 30 different cells at metaphase stage. The relative concentrations of probe DNA and blocking DNA were adjusted for each BAC clone used as probe for FISH. When BAC3 containing SSR HA4222, anchored to LG 16 was used as probe, one pair of specific signals were detected (Fig. 2a). In this case a DNA concentration ratio of 1 to 20 between BAC3 and SSR-Ha785 was used. Figure 2b shows two terminal signals in two chromosomes with similar size and morphology, when BAC6 (containing SSR HA2600) was used as probe. Here, the DNA was blocked with both SSR-Ha785 (DNA concentration ratio = 1/15) and HaRep1 (DNA concentration ratio = 1/10). Regarding C-PRINS strategy, indirect C-PRINS was applied for the localization of one single SSR site in the sunflower genetic map. To this aim, oligonucleotides corresponding to the flanking regions of ORS1247 anchored to LG 17 were used as primers. Consequently, two strong signals were observed in a pair of SAT-chromosomes corresponding to ORS1247 (Fig. 2c).
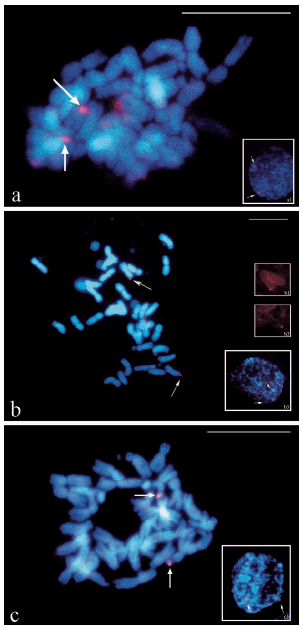
Figure 2 BAC-FISH: (a) mitotic metaphase, the arrows show hybridization signals in homologous chromosomes using BAC3 containing the SSR HA4222 (LG 16) as probe. Relation probe: DNA blocking = 1: 20. (a1) interphase nucleus showing two signals. (b) mitotic metaphase, the arrows show hybridization signals in homologous chromosomes using BAC6 containing the SSR HA2600 (LG 10) as probe. Relation probe: DNA blocking = 1: 20. (b1-b2) chromosomes staining with digoxigenin showing hybridization signals on the chromosomes (b3) interphase nucleus showing two signals. C-PRINS: (c) mitotic metaphase, the arrows show bright fluorescent signal in two chromosomes using primers of SSR ORS1247 as probe. (c1) interphase nucleus showing two signals. The counterstaining was made with DAPI. Bar =10 µm.
Discussion
In the present work, BAC-FISH and C-PRINS techniques were evaluated for the detection of single-copy DNA sequences on sunflower chromosomes, corresponding to already mapped single-site SSR markers (Kiani et al., 2007).
The detection of unique targeted sequences in plants with large genomes by FISH using probes with molecular sizes lower than 10 kbp turned out to be very difficult even after using amplification methods such as tyramide amplification (Khrustaleva and Kik, 2001; Qian and Lloyd, 2003). Thus, the use of large insert size of BAC clones turned out as an attractive strategy for physical mapping of single copy sequences on different plant species (Lapitan et al., 1997; Wang and Chen, 2005), although it requires the identification of suitable BAC specific clones.
In this work, the use of pools of overgo probes to identify BAC clones containing target single-site SSRs was more efficient than a traditional screening using specific individual probes. Through this technique, six BAC clones containing five markers of unique location, previously anchored to three linkage groups of interest: LG 10 (HA928, HA2600), LG 16 (HA4222) and LG 9 (HA2063, ORS805) were identified. These LGs encompassed previously reported QTL for resistance to S. sclerotiorum (Maringolo, 2007). The BAC-FISH technique using these BAC clones as probes in in situ hybridization assays delivered a scattered pattern of hybridization which indicates the presence of repetitive elements in these clones, as confirmed by the BAC-Southern blot performed using specific repetitive DNA probes. We observed different hybridization patterns for each BAC clone as well as differences in signal intensity for the different evaluated repetitive sequences (repetitive sequences previously reported in sunflower as well as sequence with similarity to copia-like and gypsy-like retrotransposon families). These results confirm previous reports indicating the disperse distribution of sequences with similarity to retrotransposons in the sunflower genome (Natali et al., 2006; Tang et al., 2006). Due to this reason, it was necessary to block these repetitive DNA sequences to detect single copy sequences. Thus, blocking strategies were assayed for the identification of unique target sequences by BAC-FISH.
Our study showed that the BAC-FISH technique allowed the identification of specific signals in metaphasic chromosomes for SSR HA4222 marker anchored to LG 16 and for SSR HA2600 anchored in LG 10. These results confirm that BAC-FISH is a useful tool for the detection of single copy sequences in plant species with large genomes such as sunflower. However, since the presence of large amounts of repetitive elements present in BAC clones impairs the localization of unique sequences in different species (Kim et al., 2002), the blocking conditions for these sequences for each BAC-FISH assay was optimized. We determined that the sunflower repetitive sequences, SSR-Ha785 and the HaRep1, turned out to be efficient blocking DNAs suitable for BAC-FISH applications.
In parallel, C-PRINS was tested as a simpler and faster alternative to BAC-FISH for localization of single copy sequences like SSRs anchored to know LGs. Intense fluorescent signals were detected by primer extension using oligonucleotides directed to the SSR marker ORS1247 anchored to the LG 17. As expected, when primers for ORS1247 were tested, signals were observed in two homologous chromosomes.
In this work we demonstrate that indirect C-PRINS is a useful method to detect single copy sequences in sunflower. These results agree with those reported by Kubaláková et al. (2001), who found that the C-PRINS technique is a more sensitive variant of PRINS technique and more useful for the detection of low copy number sequences. C-PRINS technique appears as a valuable alternative or complementary strategy to the BAC-FISH technique for localization of single copy sequences in the sunflower genome. In comparison, C-PRINS technique has the advantage of being easy to implement, faster, cheaper and more specific avoiding the detection of repetitive DNA.
In conclusion, we found that both techniques, BAC-FISH with the use of blocking repetitive sequences and indirect C-PRINS, are suitable strategies to detect single copy sequences in sunflower. The application of these techniques to the integration of genetic and physical maps will contribute to the chromosomal location of key genes and/or QTL associated to agronomic important traits.
Acknowledgements
This work was supported by grants from the Consejo Nacional de Investigaciones Científicas y Técnicas (PIP CONICET No. 5927), the Universidad de Buenos Aires (UBACYT No. X317, UBACYT No. X178 and UBACYT No. X446), the Agencia Nacional de Promoción Científica y Tecnológica (PICT No. 30-14119) and the Instituto Nacional de Tecnología Agropecuaria (PN No. 1952, PE No. AEBIO 1433), Argentina.
Dr. L. Poggio, Dr. N. Paniego and Dr. R. Heinz are career members of the Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET, Argentina).
The authors are grateful to Diego Fink for his technical assistance in the drawing of figures.
Dr. Hopp is a career member of the Comisión de Investigaciones Científicas de la Provincia de Buenos Aires (CIC), H.E. Hopp and Dra. L. Poggio are Professors at the Facultad de Ciencias Exactas y Naturales, University of Buenos Aires (UBA).
References
1. Al-Chaarani GR, Gentzbittel L, Huang XQ, Sarrafi A (2004). Genotypic variation and identification of QTLs for agronomic traits, using AFLP and SSR markers in RILs of sunflower (Helianthus annuus L.). Theoretical Applied Genetics 109: 1353-1360. [ Links ]
2. Baack E, Whitney K, Rieseberg L (2005). Hybridization and genome size evolution: timing and magnitude of nuclear DNA content increases in Helianthus homoploid hybrid species. New Phytologist 167: 623-630. [ Links ]
3. Bouzidi MF, Franchel J, Tao Q, Stormo K, Mraz A, Nicolas P, Mouzeyar S (2006). A sunflower BAC library suitable for PCR screening and physical mapping of targeted genomic regions. Theoretical Applied Genetics 113: 81-89. [ Links ]
4. Ceccarelli M, Sarri V, Natali L, Giordani T, Cavallini A, Zuccolo A, Jurman I, Morgante M, Cionini PG (2007). Characterization of the chromosome complement of Helianthus annuus by in situ hybridization of a tandemly repeated DNA sequence. Genome 50: 429-434. [ Links ]
5. Chen MS, Presting G, Barbazuk WB, Goicoechea JL, Blackmon B, Fang GC, Kim H, Frisch D, Yu Y-S, Sun S, Higingbottom S, Phimphilai J, Phimphilai D, Thurmond S, Gaudette B, Li P, Liu JD, Hatfield J, Main D, Farrar K, Henderson C, Barnett L, Costa R, Williams B, Walser S, Atkins M, Hall C, Budiman MA, Tomkins JP, Luo MZ, Bancroft I, Salse J, Regad F, Mohapatra T, Singh NK, Tyagi AK, Soderlund C, Dean RA, Wing RA (2002). An integrated physical and genetic map of the rice genome. The Plant Cell 14: 537-545. [ Links ]
6. Danilova TA, Birchler JA (2008). Integrated cytogenetic map of mitotic metaphase chromosome 9 of maize: resolution, sensitivity, and banding paint development. Chromosoma 117: 345-356. [ Links ]
7. Desel C, Jung C, Cai DG, Kleine M, Schmidt T (2001). High-resolution mapping of YACs and the single-copy gene HS1 (pro)-1 on Beta vulgaris chromosome by multi-colour fluorescence in situ hybridization. Plant Molecular Biology 45: 113-122. [ Links ]
8. Feng J, Vick BA, Lee MK, Zhang HB, Jan CC (2006). Construction of BAC and BIBAC libraries from sunflower and identification of linkage group-specific clones by overgo hybridization. Theoretical Applied Genetic 113: 23-32. [ Links ]
9. Fernández P, Paniego N, Lew S, Hopp HE, Heinz RA (2003). Differential representation of sunflower ESTs in enriched organspecific cDNA libraries in a small scale sequencing project. BMC Genomics 4: 40. [ Links ]
10. Fusari CM, Lia VV, Hopp HE, Heinz RA, Paniego NB (2008). Identification of Single Nucleotide Polymorphisms and analysis of Linkage Disequilibrium in sunflower elite inbred lines using the candidate gene approach. BMC Plant Biology 8: 7. [ Links ]
11. Gardiner J, Schroeder S, Polacco ML, Sanchez-Villeda H, Fang Z, Morgante M, Landewe T, Fengler K, Useche F, Hanafey M, Tingey S, Chou H, Wing R, Soderlund C, Coe EH Jr. (2004). Anchoring 9,371 maize expressed sequence tagged unigenes to the bacterial artificial chromosome contig map by two-dimensional overgo hybridization. Plant Physiology 134: 1317-1326. [ Links ]
12. Gentzbittel L, Abbott A, Galaud JP, Georgi L, Fabre F, Liboz T, Alibert G (2002). A bacterial artificial chromosome (BAC) library for sunflower, and identification of clones containing genes for putative transmembrane receptors. Molecular Genetics and Genomics 266: 979-987. [ Links ]
13. Gentzbittel L, Mestries E, Mouzeyar S, Mazeyrat F, Badaoui S, Vear F, De Labrouhe DT, Nicolas P (1999) A composite map of expressed sequences and phenotypic traits of the sunflower Helianthus annuus L. genome. Theoretical and Applied Genetics 99: 218-234. [ Links ]
14. Gosden J, Hanratty D, Starling J, Fantes J, Mitchell A, Porteous D (1991). Oligonucleotide-primed in situ DNA synthesis (PRINS): a method for chromosome mapping, banding, and investigation of sequence organization. Cytogenetics and Cell Genetics 57: 100-104. [ Links ]
15. Heesacker A, Kishore V, Gao W, Tang S, Kolkman J, Gingle A, Matvienko M, Kozik A, Michelmore R, Lai Z, Rieseberg L, Knapp S (2008). SSRs and INDELs mined from the sunflower EST database: abundance, polymorphisms, and cross-taxa utility. Theoretical and Applied Genetics 117: 1021-1029. [ Links ]
16. Islam-Faridi MN, Childs KL, Klein PE, Hodnett G, Menz MA (2002). A molecular cytogenetic map of sorghum chromosome 1: fluorescence in situ hybridization analysis with mapped bacterial artificial chromosomes. Genetics 161: 345-353. [ Links ]
17. Jiang J, Gill BS, Wang GL, Ronald PC, Ward DC (1995). Metaphase and interphase fluorescence in situ hybridization mapping of rice genome with bacterial artificial chromosomes. Proceeding of the NationalAcademy of Sciences 92: 4487-4491. [ Links ]
18. Khrustaleva L, Kik C (2001). Localization of single-copy T-DNA insertion in transgenic shallots (Allium cepa) by using ultrasensitive FISH with tyramide signal amplification. Plant Journal 25: 699-707. [ Links ]
19. Kiani SP, Talia P, Maury P, Grieu P, Heinz R, Perrault A, Nishinakamasu V, Hopp H, Gentzbittel L, Paniego N, Sarrafi A (2007). Genic analysis of plant water status and osmotic adjustment in recombinant inbred lines of sunflower under water treatments. Plant Science 172: 773-787. [ Links ]
20. Kim JS, Childs KL, Islam-Faridi MN, Menz MA, Klein RR (2002) Integrated karyotyping of sorghum by in situ hybridization of landed BACs. Genome 45: 402-412. [ Links ]
21. Koch JE, Kolvraa S, Petersen KB, Gregersen N, Bolund L (1989). Oligonucleotide-priming methods for the chromosome-specific labelling of a satellite DNA in situ. Chromosoma 98: 259-265. [ Links ]
22. Kolkman JM, Berry ST, Leon AJ, Slabaugh MB, Tang S, Gao W, Shintani DK, Burke JM, Knapp SJ (2007). Single Nucleotide Polymorphisms and Linkage Disequilibrium in Sunflower. Genetics 177: 457-468. [ Links ]
23. Koorneef M, Fransz P, de Jong H (2003). Cytogenetic tools for Arabidopsis thaliana. Chromosome Research 11: 183-194. [ Links ]
24. Kubaláková M, Vrána J, Cíhalíková J, Lysák MA, Dole J (2001). Localisation of DNA sequences on plant chromosomes using PRINS and C-PRINS. Methods in Cell Science 23: 71-82. [ Links ]
25. Kumpatla S, Mukhopadhyay S (2005). Mining and survey of simple sequence repeats in expressed sequence tags of dicotyledonous species. Genome 48: 985-998. [ Links ]
26. Lai Z, Livingstone K, Zou Y, Church SA, Knapp SJ, Andrews J, Rieseberg LH (2005) Identification and mapping of SNPs from ESTs in sunflower. Theoretical and Applied Genetics 111: 1532-1544. [ Links ]
27. Lapitan NLV, Brown SE, Kennard W, Stephens JL, Knudson DL (1997). FISH physical mapping with barley BAC clones. Plant Journal 11: 149-156. [ Links ]
28. Lee HR, Eom EM, Lim YP, Bang JW, Lee DH (2003). Construction of a garlic BAC library and chromosomal assignment of BAC clones using FISH techniques. Genome 46: 514-520. [ Links ]
29. Liu A, Burke JM (2006). Patterns of nucleotide diversity in wild and cultivated sunflower. Genetics 173: 321-330. [ Links ]
30. Maringolo CA (2007). Regiones cromosómicas asociadas a resistencia a podredumbre húmeda del capítulo de girasol (Sclerotinia sclerotiorum (lib.) de bary). Tesis de maestría Facultad de Ciencias Agrarias Universidad Nacional de Mar del Plata: 1-133. [ Links ]
31. McPherson JD, Marra M, Hillier L, Waterston RH, Chinwalla A, Wallis J, Sekhon M, Wylie K, Mardis ER, Wilson RK (2001). A physical map of the human genome. Nature Publishing Group 409: 934-941. [ Links ]
32. Menke M, Fuchs J, Schubert I (1998). A comparison of sequence resolution on plant chromosomes: PRINS versus FISH. Theoretical and Applied Genetics 97: 1314-1320. [ Links ]
33. Minelli S, Maggini F, Gelati MT, Angiolillo A, Cionini PG (2000). The chromosome complement of Olea europaea L.: characterization by differential staining of the chromatin and in-situ hybridization of highly repeated DNA sequences Chromosome Research 8: 615-619. [ Links ]
34. Natali L, Santini S, Giordani T, Minelli S, Maestrini P, Cionini PG, Cavallini A (2006). Distribution of Ty3-gypsy- and Ty1-copialike DNA sequences in the genus Helianthus and other Asteraceae. Genome 49: 64-72. [ Links ]
35. Özdemir N, Horn R, Friedt W (2004). Construction and characterization of a BAC library for sunflower (Helianthus annuus L.). Euphytica 138: 177-183. [ Links ]
36. Paniego N, Echaide M, Munoz M, Fernandez L, Torales S, Faccio P, Fuxan I, Carrera M, Zandomeni R, Suarez EY, Hopp HE (2002). Microsatellite isolation and characterization in sunflower (Helianthus annuus L.). Genome 45: 34-43. [ Links ]
37. Pashley CH, Ellis JR, McCauley DE, Burke JM (2006). EST databases as a source for molecular markers: lessons from Helianthus. Journal of Heredity 97: 381-388. [ Links ]
38. Qian X, Lloyd R (2003). Recent Developments in Signal Amplification Methods for In Situ Hybridization. Diagnostic Molecular Pathology 12: 1-13. [ Links ]
39. Sambrook J, Fritsch EF, Maniatis T (1989). Molecular Cloning: A Laboratory Manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York. [ Links ]
40. Santini S, Cavallini A, Natali L, Minelli S, Maggini F, Cionini PG (2002). Ty1 /copia- and Ty3 /gypsy-like DNA sequences in Helianthus species. Chromosoma 111: 192-200. [ Links ]
41. Schwarzacher T, Heslop-Harrison P (2000). Practical In Situ Hybridization. BIOS Scientific Publishers, Oxford, U.K. [ Links ]
42. Shizuya HB, Birren B, Kim UJ, Mancino V, Slepak T, Tachiiri Y, Simon M (1992) Cloning and stable maintenance of a 300- kilobase-pair fragment of human DNA in Escherichia coli using an F-factor based vector. Proceeding of the National Academy of Sciences 89: 8794-8797. [ Links ]
43. Tang S, Hass CG, Knapp SJ (2006). Ty3/gypsy-like retrotransposon knockout of a 2-methyl-6-phytyl-1,4-benzoquinone methyltransferase is non-lethal, uncovers a cryptic paralogous mutation, and produces novel tocopherol (vitamin E) profiles in sunflower. Theoretical and Applied Genetics 113: 783-799. [ Links ]
44. Tang S, Kishore VK, Knapp SJ (2003). PCR-multiplexes for a genome-wide framework of simple sequence repeat marker loci in cultivated sunflower. Theoretical and Applied Genetics 107: 6-19. [ Links ]
45. Tang S, Yu JK, Slabaugh B, Shintani K, Knapp J (2002). Simple sequence repeat map of the sunflower genome. Theoretical and Applied Genetics 105: 1124-1136. [ Links ]
46. Wang CJ, Chen CC (2005). Cytogenetic mapping in maize. Cytogenetic and Genome Research 109: 63-69. [ Links ]
47. Yu J-K, Tang S, Slabaugh MB, Heesacker A, Cole G, Herring M, Soper J, Han F, Chu W-C, Webb DM, Thompson L, Edwards KJ, Berry S, Leon AJ, Grondona M, Olungu C, Maes N, Knapp SJ (2003). Towards a saturated molecular genetic linkage map for cultivated sunflower. Crop Science 43: 367-387. [ Links ]
48. Yüksel B, Paterson AH (2005) Construction and characterization of a peanut HindIII BAC library. Theoretical and Applied Genetics 111: 630-639. [ Links ]
Received: August 30, 2010.
Acepted: March 28, 2011.