Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO  uBio
uBio
Compartir
Phyton (Buenos Aires)
versión On-line ISSN 1851-5657
Phyton (B. Aires) v.77 Vicente López ene./dic. 2008
ARTÍCULOS ORIGINALES
A first attempt to elucidate the amino acid sequence of some lichen lectins (With 2 Tables & 3 Figures)
El primer intento en determinar la secuencia de aminoácidos de algunas lectinas de líquenes (Con 2 Tablas y 3 Figuras)
Sacristán M, Am Millanes, B Fontaniella, Me Legaz, C Vicente
Department of Plant Physiology, The Lichen Team. Faculty of Biology, Complutense University. JoséAntonio Novais s/n, 28040 Madrid, Spain
Address Correspondence to: Prof. Dr. Carlos Vicente. Department of Plant Physiology, Faculty of Biology, José Antonio Novais st., s/n. Complutense University, 28040 Madrid, Spain. Phone number: 34-1-3944565, Fax number: 34-1-3945034. e-mail: cvicente@bio.ucm.es
Recibido/Received 10.04.2008. Aceptado/Accepted 25.06.2008.
Abstract. Secreted arginases from Evernia prunastri and Xanthoria parietina thalli, exhibiting lectin activity, have been purified to homogeneity. Analyses of both amino acid and glycoside composition are reported. Purified proteins have been subjected to tryptic digestion and some amino acid sequences have been analyzed. An undecapeptide from Evernia arginase, which shows some degree of homology with the domain to which Mn2+ binds, a heptapeptide and an undecapeptide from Xanthoria arginase have been analyzed, and homologies with some arginases from other fungi and plants have been found.
Key words: Evernia prunastri; Xanthoria parietina; Arginases; Lectins; Structure analysis.
Resumen. Arginasas segregadas por talos de Evernia prunastri y Xanthoria parietina que exhiben actividad lectina han sido purificadas a homogeneidad. Los resultados del análisis de su composición de aminoácidos y de la fracción glicídica son descriptos en este trabajo. Ambas proteínas purificadas han sido sometidas a digestión tríptica y la secuencia de aminoácidos de algunos de sus fragmentos ha sido analizada. De la arginasa de Evernia se ha identificado un undecapéptido que muestra algún grado de homología con el dominio al que se une el cofactor metálico Mn2+, mientras que de la arginasa de Xanthoria se han analizado un heptapéptido y un undecapéptido que muestran homología con arginasas de otros hongos y plantas.
Palabras clave: Evernia prunastri; Xanthoria parietina; Arginasas; Lectinas; Análisis de estructuras.
INTRODUCTION
Lichens are symbiotic associations between a fungus and a cyanobacterium (cyano-lichens) or a green alga (chlorolichens), joined to form a new biological entity different from its individual components. Recognition mechanisms used by lichens are based on the production and secretion of fungal lectins as signalling molecules (Kardish et al., 1991), which therefore develop arginase enzymatic activity (Molina et al., 1993). Whereas arginases retained on the cell wall of fungal hyphae serve for adhering algal cells (contact lectins), secreted enzymes produce adhesion of free-living algal cells (recruitment lectins), which grow near the fungal partner. Both recruitment and contact require the occurrence of receptor sites (lectin ligand) on the surface of the potential photobions, to which the lectin molecule binds. This receptor has been defined as a cell wall-pelletable, glycosylated urease in Evernia prunastri and Xanthoria parietina photobionts. Lack of the adequate lectin ligand makes incompatible the photobiont. Consequently, the lectin enters the algal cell, the concentration of polyamines increases (Molina & Vicente, 1995), and these polyamines, mainly putrescine, induce ultrastructure damages such as chloroplast disorganization, hydrolysis of the cell wall and, finally, cell death (Molina et al., 1998). Increases in polyamine concentration are the result of the arginase activity of the signaling protein. On the contrary, the occurrence of the ligand at the algae surface retains the lectin outside the cell, and preserves both the structure and function of the recognized photobiont (Legaz et al., 2004).
Previous results (Molina & Vicente, 2000; Legaz et al., 2001) indicated some similarities between secreted arginases from E. prunastri (SAE) and X. parietina (SAX) with respect to their amino acid composition (Table 1).
Table 1. Quantitative composition in amino acids (mol/mol) of Evernia prunastri (SAE) and Xanthoria parietina (SAX) according to Legaz et al. (2001) and Molina& Vicente (2000), respectively.
Tabla 1. Composición cuantitativa de aminoácidos (mol/mol) de Evernia prunastri (SAE) y Xanthoria parietina (SAX) siguiendo a Legaz et al. (2001) y Molina & Vicente (2000), respectivamente.
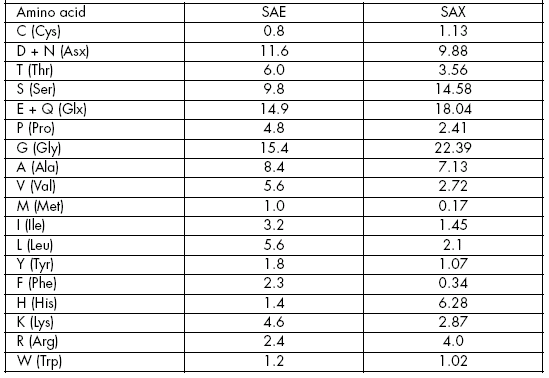
These similarities were:
Both proteins contained Asx, Cys and Trp in a similar proportion.
Content of Ala was identical for both arginases.
Although Gly and Glu were the most abundant amino acids for both arginases, their percentage value was greater in SAX than in SAE.
The lowest content of Val was found in SAX.
SAX contained lower amounts of Ile, Leu and Lys, and higher amounts of His, than SAE.
Substantial differences in the amino acid composition of the purified lectins were expected for SAE, SAX, and Dictyonema glabratum (Atheliaceae) on the basis of their phylogenetical distance (Elifio et al., 2000). This was because of (1) the presence of the amino acid Phe, (2) the fact that the content of Gly and Ser was lower than that found for SAE and SAX and, (3) the absence of Tryp. Only the low amount of Met in D. labratum lectin was similar to that found in SAX.
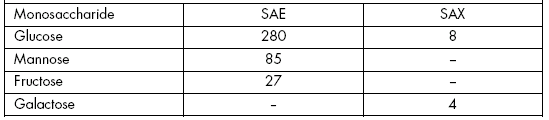
The glycosidic moiety of SAX contained equimolar amounts of both D-galactose and D-glucose (Molina & Vicente, 1996), whereas that of SAE contained 280 residues of glucose, 85 of mannose and 27 of fructose per molecule of protein (Planelles & Legaz, 1987). The lectin of D. glabratum contained mainly galactose and xylose and minor amounts of glucose, mannose, glucosamine and galactosamine (Elifio et al., 2000).
Binding of fungal arginase to glycosylated urease in the algal cell wall implies an affinity mechanism between some specific amino acid residues of the lectin and specific sugar residues of the glycoside moiety of the ligand (Table 2). This is a polygalactose chain attached to the urease polypeptide in Evernia prunastri (Pérez-Urria & Vicente, 1989). This polysaccharide moiety results to be an α- (1,4) polygalactoside, as found by Sacristán et al. (2006). Nevertheless, other receptors exist different from α-(1,4) polygalactosyl urease. This is because concanavalin A, the lectin from Canavalia ensiformis, is able to bind to the cell wall of algal cells recently isolated from Evernia prunastri and Xanthoria parietina thalli (Fontaniella et al., 2004). Concanavalin A of C. ensiformi s also develops arginase activity. This binding involves a ligand, probably a glycoprotein containing mannose, which can be isolated by affinity chromatography. Concanavalin A also brinds to the cell wall of Nosotoc from several Peltigera species (Marx & Peveling, 1983). In addition, a new lectin has been isolated from an aqueous extract of the symbiotic phenotype of Dictyonema glabratum and its cyanobacterial photobiont Scytonema sp. (Elifio et al., 2000). The lectin is a glycoprotein with a low degree of glycosylation, with both N- and O-linkages. The receptor the lectin binds to contains N-acetylgalactosamine as the most active monosaccharide for affinity relationship.
Table 2. Monosaccharide residues per molecule of secreted arginases purified from E. prunastri (SAE) and X. parietina (SAX).
Tabla 2. Residuos de monosacáridos por molécula de argininas segregadas, purificadas de E. prunastri (SAE) y X. parietina (SAX).
In this work, we analyzed the amino acid sequence of secreted arginases from E. prunastri and X. parietina to find homologies that support the function of both lectins produced by two phylogenetically unrelated lichen species.
MATERIALS AND METHODS
Plant material and chemicals
Evernia prunastri (L.) Ach. (Parmeliaceae) containing Trebouxia excen-trica as a photobiont, and Xanthoria parietina (L.) Th.Fr. (Teloschistaceae) containing Pseudotrebouxia aggregata as a photobiont, were collected during January 2003 in La Quinta (El Pardo, Madrid, Spain). Both species grow on Quercus rotundifolia Willd. They were dried under air flow, and stored in the dark for no longer than two weeks. All chemicals used in this work were from Sigma Chemical Company (St.Louis, MO, USA) unless stated otherwise.
Purification procedures
Secreted arginase from Evernia (SAE) was purified according to Legaz et al. (2001). Samples of 50 g of air-dried thalli of E. prunastri were floated on 1250 ml of buffer (10 mM Tris-HCl, pH 9.15) containing 40 mM L-arginine for 8 h at 26 °C in the dark. Then, the medium was filtered through Whatman 3MM filter paper ( W. and R. Balston Ltd, Maidstone, UK), concentrated under air flow up to 300 ml, filtered through Millipore GS filters (0.22 mm diameter, Millipore Corporation, Billerica, MA, USA) and immediately precipitated with ammonium sulphate at 50% (w/v). The mixture was stored for 2 h at 4 °C, thereafter centrifuged for 1h at 43,000 x g at 2 °C and subjected to dialysis. Protein in the dialysate was then adsorbed on calcium phosphate gel. The eluate from 0.14 M Tris-HCl buffer (pH 9.1) contained the highest arginase activity. This eluate was chromatofocused by using a LKB 8100 column (LKB, Bromma, Sweden). The pH gradient was made with 1% (w/v) ampholines (Servalyt, Serva Feinbiochemica Co., Heidelberg, Germany), pH 3.5-10, and the density gradient was prepared from 5 to 50% sucrose solutions. The electrode solutions were 1.0 M NaOH at the cathode, and 1.0 M H3 PO4 at the anode. The cooling temperature was 4 °C Fractions of 2.0 ml were collected and assayed for arginase activ-ity and pH value. The fraction eluted at pH 5.6 contained the highest arginase activity.
Alternatively, samples of 15 g of X. parietina thalli were floated on 150 ml 10 mM Tris-HCl buffer (pH 9.1) for 1h at 26 °C in the dark. Secreted arginase (SAX) was purified from the incubation media according to Molina & Vicente (2000). This was made by precipitation with ammonium sulphate at 50% saturation, adsorption on calcium phosphate gel (SAX was desorbed with 220 mM Tris-HCl) and filtration through a column of Sephadex (Pharmacia, Uppsala, Sweden) G-150 (30 cm x 3.0 cm i.d.). To test the homogeneity of the protein, 10 μg of SAE and 6.0 μg SAX in 150 μl were mixed with 75 μl glycerol. This solution was applied onto 12% polyacrylamide gels, and subjected to standard electrophoresis, employing 50 mM Tris-glycine (pH 8.3) as running buffer. The mixture SeeBlue 2 plus (Invitrogen, Carlsbad, CA, USA) was used as molecular mass marker.
SDS-PAGE
Both proteins (SAE and SAX), purified at homogeinity, were analyzed by SDS-PAGE according to Laemmli (1970). Proteins were redissolved in Laemmli buffer containing 50 μM 2-mercaptoethanol. Samples were clarified by centrifugation at 20,000 x g during 15 min at 2 °C. Supernatants were boiled for 5 min and loaded onto 15% (w/v) polyacrylamide gel and developed by SDS/PAGE. Fifty milimols Tris/0.1 M Gly containing 0.1% (w/v) SDS was used as running buffer. Protein was revealed with colloidal Coomassie blue (Gel Code Blue stain reagent, Pierce, Rockford, IL, USA), the gel was dried in vacuum and scanned. Colloidal dye was selected since its sensitivity in revealing protein bands is similar to that obtained with silver stain (Neuhoff et al., 1988), about 6.0 ng protein (2.5 ng for the silver method using bovine serum albumin). The mixture SeeBlue 2 plus, from Novex Innogenetics was used as molecular marker.
Enzymatic digestion and sequence analysis
Individual bands of SAE and SAX from polyacrylamide gels were digested with trypsine, analysing then extracted peptides in a MS spectrometer 4700 Reflector Spectrometer (Thermo) in tandem type MALDIT OF/TOF. From a first analysis, peptidic treads, after filtering the results to remove peaks corresponding to trypsine, were used to achieve a screening for peptide identification using MASCOT as a screening motor and all trypsine digested of protein stored in the SWISSPROT database. A peptide of molecular mass 1224.62, obtained from SAE and those of molecular masses 927.5 and 1478.78 from SAX were used to obtain a probable sequence de novo using a DENOVO TOOL program from Applied Biosystems.
RESULTS AND DISCUSSION
Tryptic digestion of individual bands of SAE and SAX in SDS-PAGE (Fig.1) produced different peptides. Their sequence was analyzed as described in methods. The sequence of the fragment of a molecular mass value of 1224.62, obtained after tryptic digestion of SAE, was revealed as PPGGHNKAKYR (Pro-Pro-Gly-Gly-His-Asn-Lys-Ala-Lys-Tyr-Arg). The tripeptide PPG also occurred in arginases from Arabidopsis thaliana (Perozich et al., 1998), Schizosaccharomyces pombe (Van Huffel et al., 1994) and Pinus taeda (Todd & Gifford, 2003), whereas GHN was repeated in arginases from A. thaliana and G. max, KAK in those from Saccharomyces cerevisiae (Kawamoto et al., 1987), and GGH in those from P. taeda (Fig. 2 and 3).
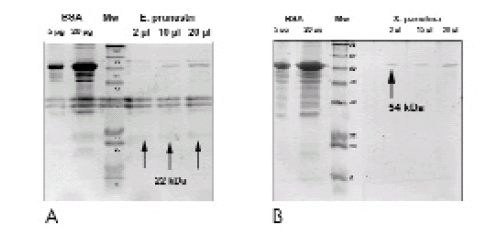
Fig 1. SDS-PAGE of both SAE (A), which produces a band of about 22 kDa, and SAX (B), that produces a band of about 54 kDa, previously dialyzed against 10mM Tris-HCl buffer (pH 9.1). SDS-PAGE was achieved in parallel to two different amounts of bovine serum albumin (BSA) as a control, and the mixture of molecular markers (Mw).
Fig 1. SDS-PAGE de SAE (A), que produce una banda de aproximadamente 22 kDa, y SAX (B), que produce una banda de aproximadamente 54 kDa, previamente dializadas en un buffer de 10mM Tris-HCl (pH 9,1). SDS-PAGE se obtuvo paralelamente a dos cantidades diferentes de albúmina de suero bovino (BSA) que se usaron como control, y la mezcla de marcadores moleculares (Mw).
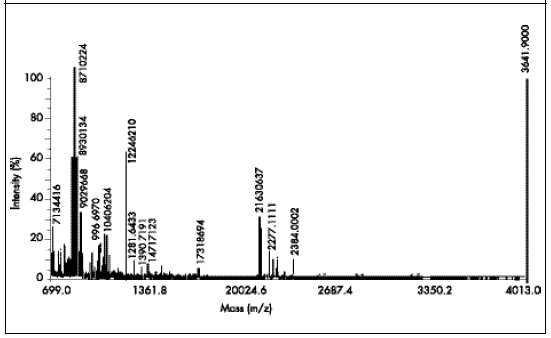
Fig 2. Peptidic thread of SAE used to select the significant peak of 1224.62 Da to be sequenced.
Fig 2. Enlaces peptídicos de SAE utilizados para seleccionar el pico significativo de1224,62 Da que va ser secuenciado.
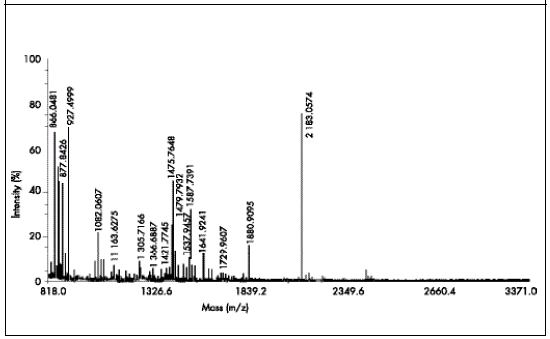
Fig 3. Peptidic thread of SAE used to select the significant peak of 1479.78 Da to be sequenced.
Fig 3. Enlaces peptídicos de SAE utilizado para seleccionar el pico significativo de 1479,78 Da que va a ser secuenciado.
Tryptic digestion of SAX produced two different peptides. One of them, with a molecular mass of 927.5, had a sequence YLYELAR (Tyr-Leu-Tyr-Glu-Leu-Ala-Arg) in MS-MS. The tripeptide ELA was repeated in the sequences of A. thaliana, S. pombe and G. max. The second fragment, with a molecular mass of 1479.78 and a sequence KNGLKFGTSTR (Lys-Asn-Gly-Leu-Lys-Phe-Gly-Thr-Ser-Thr-Arg), contained the tripeptides TST and KFG, repeated in arginase from G.max and KGN, repeated in arginase from Agaricus bisporus (Wagemaker et al., 2005). No homologies were found between SAE and SAX with arginase from Neurospora crassa (Marathe et al., 1998).
It has been proposed that His141 of rat liver arginase directly binds arginine, acting as a catalytic proton shuttle (Canales et al., 2001). Glu277 is also involved in arginine binding, which implies the corrected orientation of the substrate for the nucleophilic attack of Mn2+-bridging water (Kanyo et al., 1996). Very similar conformation can be found in the active center of yeast arginase (Sumrada & Cooper, 1984), and soybean arginase (Goldraij et al., 1998). In soybean arginase, the tripeptide Gly-His-Asn, which contains the active rest of His for the association to the substrate, is clearly repeated in the undecapeptide obtained from SAE. However, Tyr rest found in the heptapeptide obtained from SAX did not have any equivalence to Tyr rests involved in the site for Ca2+ binding, required to develop lectin activity of secreted arginases in the recognition of compatible photobionts (Legaz et al., 2004). A motif Tyr100, Asp208 and Arg228 in the well-known legume lectin, concanavalin A, is required to Ca2+ binding (Shoham et al., 1979) and repeated in Tyr173, Asp280 and Arg301 from soybean arginase (Goldraij et al., 1998). Curiously, concanavalin A also binds to mannose-containing ligands in the cell wall of lichen photobionts, as it has been found by Fontaniella et al. (2004).
Protein samples (700 pmol) were applied to an amino acid sequencer, but an N-terminal sequence could not be assigned, suggesting that the N-terminal amino acid is possibly blocked and involved in the glycosylation of the protein.
ACKNOWLEDGEMENTS
This work was supported by a grant from the Ministerio de Ciencia y Tecnología (Spain), BFI2003-06234. We wish to thank Miss Raquel Alonso for her excellent technical assistance.
REFERENCES
1. Canales, M., L. Westermeyer & N. Carvajal (2001). Molecular dymanics simulations of active site mutants of rat liver arginase. Electronic Journal of Biotechnology 4: 153-159. [ Links ]
2. Elifio, S.L., M.L.C.C. Silva, M. Iacomini & P.A.J. Gorin (2000). A lectin from the lichenized Basidiomycete Dictyonema glabratum. New Phytologist 148: 327-334. [ Links ]
3. Fontaniella, B., A.M. Millanes, C. Vicente & M.E. Legaz (2004). Concanavalin A binds to mannose-containing ligand in the cell wall of some lichen photobionts. Plant Physiology and Biochemistry 42: 773-779. [ Links ]
4. Kang, J.H. & Y.D. Cho (1990). Purification and properties of arginase from soybean, Glycine max, axes. Plant Physiology 93: 1230-1234. [ Links ]
5. Kanyo, Z.F., L.R. Scolnick, D.E. Ash & D.W. Christianson (1996). Structure of a unique binuclear manganese cluster in arginase. Nature 383: 554-557. [ Links ]
6. Kardish, N., L. Silberstein, N. Flemminger & M. Galun (1991). Lectin from the lichen Nephroma laevigatum. Localization and function. Symbiosis 11: 47-62. [ Links ]
7. Kawamoto, S., Y. Amaya, K. Murakami, F. Tokunaga, S. Iwanaga, K. Kobayashi, T. Saheki, S. Kimura & M. Mori (1987). Complete nucleotide sequence of cDNA and deduced amino acid sequence of rat liver arginase. Journal of Biological Chemistry 262: 6280-6283. [ Links ]
8. Laemmli, U.K. (1970). Cleavage of structural proteins during the assembly of the head of bacterio-phage T4. Nature 227: 680-685. [ Links ]
9. Legaz, M.E., B. Fontaniella, A.M. Millanes & C. Vicente (2004). Secreted arginases from phylogenetically far-related lichens species act as cross-recognition factors for two different algal cells. European Journal of Cell Biology 83: 435-446. [ Links ]
10. Legaz, M.E., C. Vicente & M.M. Pedrosa (2001). Binding of lichen phenolics to purified secreted arginase from the lichen Evernia prunastri. Journal of Biochemical and Molecular Biology 34: 194-200. [ Links ]
11. Marathe, S., Y.G. Yu , G.E. Turner, C. Palmier & R.L. Weiss (1998). Multiple forms of arginase are differentially expressed from a single locus in Neurospora crassa. Journal of Biological Chemistry 273: 29776-29785. [ Links ]
12. Marx, M. & E. Peveling (1983). Surface receptors in lichen symbionts visualized by fluorescence microscopy after use of lectins. Protoplasma 114: 52-61. [ Links ]
13. Molina, M.C., E. Muñiz & C. Vicente (1993). Enzymatic activities of algal-binding protein and its algal cell wall receptor in the lichen Xanthoria parietina. An approach to the parasitic basis of mutualism. Plant Physiology and Biochemistry 31: 131-142. [ Links ]
14. Molina, M.C., E. Stocker-Wörgötter, R. Turk, C. Bajon & C. Vicente (1998). Secreted, glycosylated arginase from Xanthoria parietina thallus induces loss of cytoplasmic material from Xanthoria photobionts. Cell Adhesion and Communication 6: 481-490. [ Links ]
15. Molina, M.C. & C. Vicente (1995). Correlationships between enzymatic activity of lectins, putrescine content and chloroplast damage in Xanthoria parietina phycobionts. Cell Adhesion and Communication 3: 1-12. [ Links ]
16. Molina, M.C. & C. Vicente (1996). High-performance liquid chromatographic characterization of two lichen lectins with arginase activity differing in their glycosyl moiety. Journal of Liquid Chromatography and Related Techniques 19: 2101-2115. [ Links ]
17. Molina, M.C. & C. Vicente (2000). Purification and characterization of two isolectins with arginase activity from the lichen Xanthoria parietina. Journal of Biochemistry and Molecular Biology 33: 300-307. [ Links ]
18. Neuhoff, V. , N. Arold, D. Taube & W. Ehrhardt (1988). Improved staining of proteins in polyacry-lamide gels including isoelectric focusing gels with clear background at nanogram sensi-tivity using Coomassie Brilliant Blue G-250 and R-250. Electrophoresis 9: 255-262. [ Links ]
19. Pérez-Urria, E. & C. Vicente (1989). Purification and some properties of a secreted urease from Evernia prunastri. Journal of Plant Physiology 133: 692-695. [ Links ]
20. Perozich, J., J. Hempel & S.M. Morris (1998). Roles of conserved residues in the arginase family. Biochimica Biophysica Acta 1382: 23-37. [ Links ]
21. Planelles, V. & M.E. Legaz (1987). Purification and some properties of the secreted arginase of the lichen Evernia prunastri and its regulation by usnic acid. Plant Science 51: 9-16. [ Links ]
22. Sacristán, M., A.M. Millanes, M.E. Legaz & C. Vicente (2006). A lichen lectin specifically binds to the a-1,4-polygalactoside moiety of urease located in the cell wall of homologous algae. Plant Signaling & Behavior 1: 23-27. [ Links ]
23. Shoham, M., A. Yonath, J.L. Sussmann, J. Moult, W. Traub & A.J. Kalb (1979). Crystal structure of demetallized concanavalin A: the metal binding region. Journal of Molecular Biology 131: 137-155. [ Links ]
24. Sumrada, R.A. & T.G. Cooper (1984). Nucleotide sequence of the Saccharomyces cerevisiae arginase gene (Car1) and its transcription under various physiological conditions. Journal of Bacteriology 160: 1078-1087. [ Links ]
25. Todd, C.D. & D.J. Gifford (2003). Loblolly pine arginase responds to arginine in vitro. Planta 217: 610-6155. [ Links ]
26. Van Huffel, C., E. Dubois & F. Messenguy (1994). Cloning and sequencing of Schizosaccharomyces pombe car1 gene encoding arginase. Expression of the arginine anabolic and catabolic genes in response to arginine and related metabolites. Yeast 10: 923-933. [ Links ]
27. Wagemaker, M.J., W. Welboren, C. van der Drift, M.S. Jetten, L.J. Va n Griensven & H.J. Op den Camp (2005). The ornithine cycle enzyme arginase from Agaricus bisporus and its role in urea accumulation in fruit bodies. Biochem. Biophys. Acta 1681: 107-115. [ Links ]














