Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Phyton (Buenos Aires)
On-line version ISSN 1851-5657
Phyton (B. Aires) vol.79 no.2 Vicente López July/Dec. 2010
ARTÍCULOS ORIGINALES
Comparison of isozyme transformation in maize as a result of insertion of the chitinase gene
Comparación de la transformación de isoenzimas en maíz como resultado de la inserción del gen de la quitinasa
Yan PM1, HF Zhang1, Q Wang1, XY Yan1, Y Sun2
1 Taiyuan Normal University, Shanxi, Taiyuan 030012, P. R. China.
2 Agricultural Biotechnolgy Center of Shanxi, Taiyuan 030031, China.
Address Correspondence to: Dr. Yan Pingmei. Taiyuan Normal University, Shanxi, Taiyuan 030012, P. R. China. e-mail: yanpingmei2010@gmail.com
Recibido- Received 15.VII.2010.
Aceptado - Accepted 25.VIII.2010.
Abstract. Isozymes of peroxidase (POD), catalase (CAT), esterase (EST) and superoxide dismutase (SOD) were analyzed on transgenic maize (with external chitinase gene) and its parent by vertical polyacrylamide gel electrophoresis (PAGE). This study was made using shoots at the fourth leaf stage. Results showed that: POD and EST were detected in 6 bands. POD-2 and POD-3 were present at the bud and seedling stages. POD-1, POD-4, POD-5 and POD-6 were only present at the seedling stage. POD-6 expressed stronger in the transgenic maize with chitinase than in its parent. EST-2 was present only at the bud stage, and its expression in transgenic maize was stronger than that in its parent. EST-5 only existed at the seedling stage. EST-4 did not exist in the parent maize seedlings and EST-1, EST-3 and EST-6 were present at the bud or seedling stage. Four bands were detected for CAT. CAT-1 and CAT-3 were weaker bands than the others. CAT-3 in transgenic maize was stronger than in its parent. Three bands of SOD were detected; SOD-1 and SOD-2 existed at the bud and seedling stages, but SOD-3 was not shown in buds of the parent corn. All data showed that the expression of isozymes in transgenic and parent maize had obvious differences.
Keywords: Chitinase; Isozyme; Maize; Antioxidant enzyme system.
Abbreviations: POD: peroxidase; CAT: catalase; EST: esterase; SOD: superoxide dismutase; PAGE: polyacrylamide gel electrophoresis; SDS: sodium lauryl sulphate; TEMED: tetra-methylethylenediamine.
Resumen. Las isoenzimas de la peroxidasa (POD), catalasa (CAT), estearasa (EST) y superóxido dimutasa (SOD) fueron analizadas en maíz transgénico (con el gen externo de la quitinasa) y su progenitor por electoforesis de gel de poliacrilamida (PAGE). Este estudio fue hecho utilizando tallos en el estado fenológico de cuarta hoja. Los resultados mostraron que: POD y EST se detectaron en 6 bandas. POD-2 y POD-3 estuvieron presentes en los estados fenológicos de yema y plántula. POD-1, POD-4, POD-5 y POD-6 sólo estuvieron presentes en el estado de plántula. POD-6 se expresó con más firmeza en el maíz transgénico con quitinasa que en su progenitor. EST-2 estuvo presente sólo en el estado de yema, y su expresión en el maíz transgénico fue más firme que en su progenitor. EST-5 sólo existió en el estado de plántula. EST-4 no existió en las plántulas progenitoras de maíz, y EST-1, EST-3 y EST-6 estuvieron presentes en los estados de yema o plántula. Se detectaron cuatro bandas para CAT. Las bandas fueron más débiles para CAT-1 y CAT-3. Este último se expresó más firmemente en maíz transgénico que en su progenitor. Se detectaron tres bandas de SOD: SOD-1 y SOD-2 existieron en los estados de yema y plántula, pero SOD-3 no estuvo presente en las yemas del maíz progenitor. Todos los datos mostraron que la expresión de las isoenzimas tuvieron diferencias obvias en maíz transgénico versus su progenitor.
Palabras clave: Quitinasa; Isoenzimas; Maíz; Sistema enzimático antioxidante.
Abreviaturas: POD: peroxidasa; CAT: catalasa; EST: estearasa; SOD: superóxido dismutasa; PAGE: electroforesis en gel de poliacrilamida; SDS: dodecil sulfato de sodio; TEMED: tetra-metiletilendiamina.
INTRODUCTION
Consumers pay more attention to the availability of transgenic plants in China as a result of the rapid development and importation of these kinds of plants in this country (Rui et al., 2005; Rui et al., 2006). With the introduction of exogenous genes, old genomes and metabolic balances could be broken in receptor plants. This could result in physiological and/or biochemical changes (e.g., changes in enzyme isozymes and nutrition).
Chitinase exists extensively in animals, plants and microbes. It may constitute an induced defense mechanism in some plants. Its distribution, characteristics, localization, induction and presence in transgenic plants have been investigated (Lan & Chen, 1998). It has been an effective approach to prevent plant, insect and fungi diseases (Ouyang et al., 2001). Maize is the second cereal crop in China, and most maize diseases come from fungi. This makes meaningful to breed transgenic maize with the chitinase gene.
Isolation of isozymes has become an important technique in molecular biology to conduct research on biological evolution, physiology and biochemistry, taxonomy, and genetic science (Rider & Taylor, 1980; Luo et al., 1999). Today, control of the insertion site for exogenous genes is difficult, and little is known about this subject. Insertion of exogenous genes could change content of plant metabolic substances and enzymes, which are the primary cause of transgenic plant safety (Zhao, 1994). In this study, transgenic maize seeds (with the chitinase gene) were used to (1) study changes of isozymes in the antioxidant enzyme system [POD (peroxidase), CAT (catalase), SOD (Superoxide Dismutase) and EST (esterase)] and (2) provide a basis for transgenic plant ecological safety.
MATERIALS AND METHODS
Plant materials. The transgenic and non-transgenic maize seeds were provided by the Agricultural Biotechnology Center of Shanxi, Taiyuan, China. The transgenic maize was achieved by the pollen-mediated approach on maize (Zea mays L.) inbred with line Hai 921. Plasmid DNA of pGL C -RC-1 was mixed with fresh pollen of maize inbreds in sucrose solution (Wang et al., 2001). Plump and smooth seeds were selected for research.
Culture of maize. Plump and healthy seeds were soaked in water during 24 hours, and placed on white porcelain plates. During the first three days, seedlings were cultured in distilled water, and thereafter cultured in 1/2 Hoogland solution (Table 1). Juveniles were transplanted to light loam soil after root length was 2 cm. Samplings were conducted after sprouting and at the four-leaf-stage.
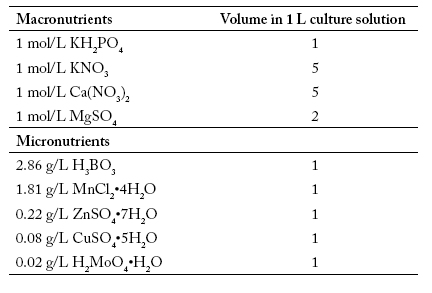
Table 1. Composition of culture solution.
Tabla 1. Composición de la solución de cultivo.
Experimental methods
Discontinuous polyacrylamide gel electrophoresis
The enzyme extracts underwent SDS-PAGE according to Sanchez' method (Sanchez et al., 2003) on vertical slab gels. The polyacrylamide gel consisted of a (1) 4.5%T/1.2%Cbis stacking gel in 0.125 M Tris-HCl buffer (pH 6.8) with 0.1% (w/v) SDS, and (2) 12.5%T/3.3%Cbis resolving gel in 0.375 M Tris-HCl buffer (pH 8.8) with 0.1% (w/v) SDS.
Thereafter, we mixed 100μL enzyme solution with 10μL 0.025% bromophenol blue. Samples of 30-40μL were added into the sample with Syringes. The voltage was regulated to 150V, stabilizing it to 200V after bromophenol blue come into a separation gel. The electrophoresis was ended when the distance from bromophenol blue to bottom was 1 cm (Yang & Zeng, 1984).
Isozyme of POD
Sample preparation. Fresh maize shoots were washed by diluted water and dried using filter paper. One gram of sample was ground in a precooled mortar with 5 ml of 1mol/L phosphate buffer (pH 7.8). Materials were stored at 4 °C in the refrigerator for 2 h. Thereafter, they were centrifuged for 10 min at 10000 rpm. The extracted material contained the enzyme sample, which was stored at -20 °C.
Staining - Acetic acid - benzidine method (Hu & Wan, 1985). The acetic acid - benzidine solution was prepared with 2 g benzidine, 18 ml Acetic acid and 72 ml double distilled water. The staining solution was made using 5 ml Acetic acid - benzidine solution, 2 ml 3% H2O2 and 93 ml double distilled water. The gel was washed three times using double distilled water after electrophoresis. It was then soaked 5-10 min in the staining solution, and photographs were taken after bands were clear.
Isozyme of CAT
Preparation of sample was similar to that of POD. However, concentration of separation gel was 10-11%. Also, 0.5% soluble starch solution was added to the separation gel.
Staining. It was made using two solutions: A and B.
A Solution: 25 ml 3% H2O2, 5 ml 0.1mol/L phosphate buffer (pH 7.0) and 3.5 ml 0.1mol/L Na2S2O3 .
B Solution: 25 ml 0.09mol/L KI were dissolved into 25 ml double distilled water.
The gel was soaked in the A solution for 15 min at room temperature, washed with double distilled water, and placed into the B solution. The resulting material was fixed in 10% glycerin.
Isozyme of EST
Sample preparation. The operational procedures were the same as for POD.
Staining solution preparation (Rider & Taylor, 1980): 50 mg α- naphthylacetate, 50 mg β- naphthylacetate and 100 mg fast blue RR salt were dissolved in 2 ml acetone solution (Vacetone/Vwater=1:1). The resulting solution was diluted with 150 ml 0.1mol/L phosphate buffer (pH 6.5). The gel was washed with double distilled water after electrophoresis was finished. This material was then put into the staining solution for 1-2 h at 25 °C, soaked with 7% acetic acid solution, and photographs were then taken.
Isozyme of SOD
Sample preparation. Fresh maize shoots were washed with distilled water and dried with filter paper. One gram of sample was ground in a precooled mortar with 1.5 ml 0.05 mol/L phosphate buffer (pH 7.8). The resulting material was centrifuged for 40 min at 12000 rpm at 4 °C. One milliliter 0.05 mol/L phosphate buffer (pH 7.8) was added to the precipitate. It was then ground and centrifuged for 40 min at 12000 rpm at 4 °C. This allowed us to obtain the enzyme sample.
Staining. Staining solution preparation (Wang, 1996): Nitroblue tetrazolium (NBT) 25 μmol/L, lactochrome 0.01%, 50 mmol/L phosphate buffer (pH 7.8), containing 1mmol/L EDTA. The gel was placed into 80ml NBT nitroblue tetrazolium solution for 15 min. It was then soaked in lactochrome solution for 5 min, and thereafter immersed in 50 mmol/L phosphate buffer (pH 7.8) containing 1 mmol/L EDTA. The gel was illuminated at 10 cm height using 40 W fluorescent lamps until transparent bands emerged.
RESULTS
Comparison of POD isozyme of transgenic maize and its parent. Results showed that POD isozyme of transgenic maize and non-transgenic maize have six bands (POD-1, POD-2, POD-3, POD-4, POD-5 and POD-6). However, the expression quantity of each band was different in transgenic maize from that of non-transgenic maize at the bud and seedling stages. POD-1, POD-4, POD-5 and POD-6 appeared only at the seedling stage, but they did not exist at the bud stage. Transgenic maize expressed less POD-5 and more POD-6 than non-transgenic maize (Fig. 1).

Fig. 1. POD differentiation in different tissues of corn genotypes. Gel at the left is at the bud stage. Gel at the right is at the seedling stage.
TR: transgenic maize; CK: Control. Every band represents an isozyme.
Fig. 1. Diferenciación de POD en diferentes tejidos de genotipos demaíz. El gel a la izquierda corresponde al estado de yema, mientras que el de la derecha corresponde al estado de plántula.
TR: maíz transgénico; CK: Control. Cada banda representa una isoenzima.
Comparison of CAT isozyme in transgenic maize versus its parent. Results showed that CAT isozyme had four bands in transgenic and non-transgenic corns: CAT-1, CAT-2, CAT-3 and CAT-4. CAT-1 and CAT-3 expressed less than CAT-2 and CAT-4, and they existed only at the seedling stage; transgenic maize expressed more CAT-1 and CAT-3 than non-transgenic maize. CAT-4 was the primary band of CAT isozyme; there was no difference between transgenic maize and its parent on this regard (Fig. 2).
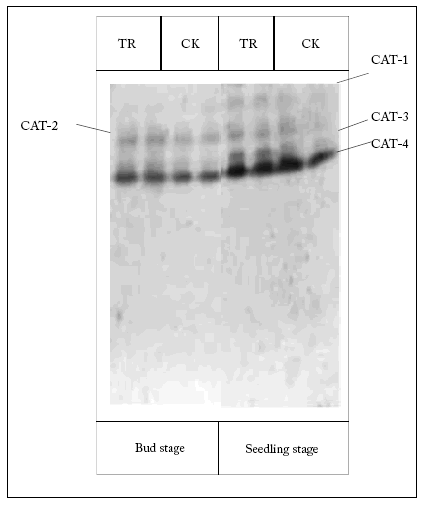
Fig. 2. Differentiation of CAT in different tissues of different corns. The left gel is at the bud stage, and the right gel is at the seedling stage.
TR: transgenic maize; CK: control. Every band represents an isozyme.
Fig. 2. Diferenciación de CAT en diferentes tejidos de genotipos de maíz. El gel a la izquierda corresponde al estado de yema, mientras que el de la derecha corresponde al estado de plántula.
TR: maíz transgénico; CK: Control. Cada banda representa una isoenzima.
Comparison of EST isozyme between transgenic maize and its parent. EST isozyme of transgenic maize showed six bands: EST-1, EST-2, EST-3, EST-4, EST-5 and EST-6; non-transgenic maize only expressed five bands: EST-1, EST-2, EST-3, EST-5 and EST-6 (Fig. 3). EST-2 only existed at the bud stage, and its expression quantity in transgenic maize was more than that in non-transgenic maize. Transgenic maize expressed EST-4 at the seedling stage, but nontransgenic maize did not.

Fig. 3. Differentiation of EST in different tissues of different corns. The left gel is at the bud stage, and the right gel is at the seedling stage.
TR: transgenic maize; CK: control. Every band represents an isozyme.
Fig. 3. Diferenciación de EST en diferentes tejidos de genotipos de maíz. El gel a la izquierda corresponde al estado de yema, mientras que el de la derecha corresponde al estado de plántula.
TR: maíz transgénico; CK: Control. Cada banda representa una isoenzima.
Comparison of SOD isozyme between transgenic maize and its parent. SOD isozyme of transgenic maize had three bands: SOD-1, SOD-2 and SOD-3 (Fig. 4). Isozymes SOD-1 and SOD-2 existed in transgenic maize and its parent at the bud and seedling stages. However, SOD-3 was not detected at the bud stage in non-transgenic maize.

Fig. 4. Differentiation of SOD in different tissues of different corns. The left gel is at the bud stage and the right gel is at the seedling stage.
TR: transgenic maize; CK: control. Every band represents an isozyme.
Fig. 4. Diferenciación de SOD en diferentes tejidos de genotipos de maíz. El gel a la izquierda corresponde al estado de yema, mientras que el de la derecha corresponde al estado de plántula.
TR: maíz transgénico; CK: Control. Cada banda representa una isoenzima.
DISCUSSION
It is well known that phenotypic traits are the result of a series of metabolic activities, which are despondent on many enzymes; their activities and concentrations regulate the metabolic reactions. More than 50% enzymes consist of isozymes which catalyze the same reaction, but isozymes have the highest reaction speeds at different conditions, such as substrate concentration, pH value, temperature and distribution in different organs (Chen et al., 2001).
POD, CAT, EST and SOD are four kinds of antioxidant enzymes which are induced under stress conditions. When an external gene is inserted into another genome, the foreign protein of that external gene is perhaps regarded as an impurity by the inherited antioxidant systems. Therefore, it is important to study the isozyme changes in POD, CAT, EST and SOD after inserting the foreign gene. The isozymes of POD, CAT, EST and SOD in transgenic maize were obviously different from those in its parents. The number (number of bands) and concentration (color depth) of isozymes usually increased. Transgenic maize expressed more POD-6, CAT-1 and CAT-3, but less POD-5, than non-transgenic maize. Also, EST-4 and SOD-3 were expressed only in transgenic maize. This indicates that the insertion and expression of the chitinase gene changed the parent map of isozymes.
Genetic engineering could accelerate biological mutation and evolution, but insertion of exogenous genes can also alter the inherited genome. This could result into either deactivation or disorder of some enzymes. Thus, the number (number of bands) and concentration (color depth) of increasing isozymes may enhance the resistance and adaptation to stress (Chen et al., 2001).
ACKNOWLEDGEMENTS
The authors appreciate the financial support from the National Science Foundation of Shanxi Province, China (No. 2009011038-3).
REFERENCES
1. Chen, X., S. Liu, R. Zhu, T. Yu & Y. Zhu (2001). Isozyme Locus Analysis in Shoots for Different Types of Hybrids and Their Parents in Rice. Journal of Wuhan Botanical Research 19: 181-186. [ Links ]
2. Hu, N.S. & G. Wan (1985). Isozyme technology and its application. Changsha: Science and Technology Press of Hunan October, pp:2-17. [ Links ]
3. Lan, H. & Z. Chen (1998). Chitinase and its research progress. Life Science Research 2: 163-169. [ Links ]
4. Luo, J., L. Zhang & J. Yang (1999). Biochemistry. A Short Course (Third edition). Beijing: China Higher Education Press, pp: 97-98. [ Links ]
5. Ouyang, S., B. Xie, K. Zhao & L. Feng (2001). Advances on Chitinase Genes Transformation. Biotechnology Information 3: 28-29. [ Links ]
6. Rider C.C. & C.B. Taylor (1980). Isoenzymes. New York Chapman and Hall, pp: 1-78. [ Links ]
7. Rui, Y.K., G.X.Yi, J. Zhao, B.M. Wang, Z.H. Li, Z.X. Zhai, Z.P. He & Q.X. Li (2005). Changes of Bt Toxin in Rhizosphere of Transgenic Bt-Cottton and its Influence on Soil Functional Bacteria. World Journal of Microbiology and Biotechnology 21: 1279-1284. [ Links ]
8. Rui, Y.K., H.X. Zhang, J. Guo, K.L. Huang, B.H. Zhu & Y.B. Luo (2006). Heavy metal content in transgenic soybean oil from Beijing market. Agro Food Industry Hi-tech 17: 35-36. [ Links ]
9. Sanchez, I., S. Seseña & L. Palop (2003). Identification of lactic acid bacteria from spontaneous fermentation of 'Almagro' eggplants by SDS-PAGE whole cell protein fingerprinting. International Journal of Food Microbiology 82: 181-189. [ Links ]
10. Wang, Z.R. Plant Allozyme Analysis (1996). Beijing Science Press. October, pp: 95-112. [ Links ]
11. Wang, J.X., Y. Sun, G.M. Cui & J.J. Hu (2001). Transgenic Maize Plants Obtained by Pollen-mediated Transformation. Acta Botanica Sinica 43: 275-279. [ Links ]
12. Yang, T.X. & M.Q Zeng (1984). The changes of isozyme, soluble protein and the dry matter accumulation during endosperm and embryo development in maize. Acta Photophysiologica Sinica 10: 77-86. [ Links ]
13. Zhao, B.(1994). Plant gene engineering technology and transgenic plants. Journal of Zibo College 2: 66. [ Links ]














