Servicios Personalizados
Revista
Articulo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares en
SciELO
Similares en
SciELO  uBio
uBio
Compartir
Phyton (Buenos Aires)
versión On-line ISSN 1851-5657
Phyton (B. Aires) vol.81 no.1 Vicente López ene./jun. 2012
ORIGINAL ARTICLES
Bacterial diversity associated with the rhizosphere of wheat plants (Triticum aestivum): Toward a metagenomic analysis
Diversidad bacteriana asociada a la rizósfera de plantas de trigo (Triticum aestivum): Hacia un análisis metagenómico
Velázquez-Sepúlveda I, MC Orozco-Mosqueda, CM Prieto-Barajas, G Santoyo
Instituto de Investigaciones Químico Biológicas. Universidad Michoacana de San Nicolás de Hidalgo, México.
Address Correspondence to: Gustavo Santoyo. Instituto de Investigaciones Químico Biológicas. Universidad Michoacana de San Nicolás de Hidalgo. Ciudad universitaria edif. A1', 58030. Morelia, Michoacán, México, e-mail: gustavo_santoyo@yahoo.com ; gsantoyo@umich.mx
Recibido / Received 3.IX.2011.
Aceptado / Accepted 09.11.2011.
Abstract. Rhizospheric soil is one the largest reservoirs of microbial genetic diversity. Before conducting a large-scale metagenomic analysis of an environment, such as a rhizospheric soil, it is necessary to perform a pre-screening of the resident genetic diversity. In this study, we analyzed the bacterial diversity associated with the rhizosphere of wheat plants by PCR amplification, construction of a library and sequencing of 16S rDNA genes. Thirty OTUs were detected, including the Classes Alfaproteobacteria, Betaproteobacteria, Deltaproteobacteria, Gammaproteobateria, Actinobacteria, Bacilli, Clostridia and Uncultivable bacteria. Within the Gammaproteobacteria class, the genera Pseudomonas, Stenotrophomonas and Bacillus were the most abundant, since they corresponded to 40% of the whole ribosomal library. Phylogenetic analysis showed that most of the ribosomal sequences are grouped into clades that belong to common rhizospheric or bulk-soil bacteria. To determine whether the sample is significantly diverse, a Shannon-Wiener test was performed, resulting in a rate of 3.8 bits per individual. Our results suggest that the rhizosphere of wheat plants is highly diverse and results an excellent candidate for metagenomic analysis.
Keywords: Bacterial diversity; Rhizosphere; Wheat; 16S ribosomal genes.
Resumen. El suelo rizosférico es uno de los reservorios más grandes de la diversidad genética microbiana. Antes de realizar un estudio metagenómico a gran escala de un ambiente, tal como el suelo rizosférico, es necesario realizar un análisis previo de la diversidad genética residente. Por ello, en este trabajo se detectó la diversidad bacteriana asociada a la rizósfera de plantas de trigo por medio de la amplificación de PCR, construcción de una biblioteca y secuenciación de los genes 16S ribosomales. Se detectaron 30 OTUs, incluyendo las Clases Alfaproteobacteria, Betaproteobacteria, Deltaproteobacteria, Gammaproteobateria, Actinobacteria, Bacilli, Clostridia y bacterias no cultivables. Dentro de las Gammaproteobateria, los géneros más abundantes fueron Pseudomonas, Stenotrophomonas y Bacillus, ya que correspondieron al 40% del total de la biblioteca ribosomal completa. El análisis filogenético demostró que la mayoría de las secuencias ribosomales se agrupan en los clados que pertenecen a bacterias comunes del suelo o rizosféricas. Para determinar si la muestra es bastante diversa se llevó a cabo una prueba de Shannon-Wiener, resultando en una tasa de 3,8 bits por individuo. Nuestros resultados sugieren que la rizósfera de plantas de trigo es muy diversa y resulta un candidato excelente para otro tipo de análisis metagenómicos.
Palabras clave: Diversidad bacteriana; Rizósfera; Trigo; Genes 16S ribosomales.
INTRODUCTION
Wheat (Triticum aestivum) is one of the most important crops around the world, which is widely consumed by the human population. To generate a good wheat production it is common, especially in developing countries, the use of chemicals to eliminate diseases caused by various pathogens. However, it is well known the damage that can be generated in the environment and human health because of their use (He et al., 2005). Likewise, it is known that some soil microbial communities near the plant roots, an area better known as rhizosphere, may limit or inhibit the growth of potential pathogens (Weller, 1988; Ahmad et al., 2008). Also, some microorganisms and especially some bacteria, which are known as PGPR (Plant-Growth Promoting Rhizobacteria) can promote plant growth and health (Schroth & Kloepper, 1978; Hayat et al., 2010). Recent studies have shown that the use of PGPR bacteria can reduce chemical use in crops (Adesemoye et al., 2009; Adesemoye & Kloepper, 2009).
Some bacterial inhabitants of the rhizosphere can live as saprophytes, while others can penetrate the tissues of the root and form root nodules. There, bacteria known as rhizobia can fix nitrogen and convert it into ammonium, which is assimilated by the plant (Fauvart et al., 2011). Also, other species can penetrate and colonize the roots, stems, leaves and seeds. These bacteria are known as endophytes and may play important roles as promoters of plant growth and induce protective responses against plant pathogens (Ryan et al., 2008). Interestingly, it has been suggested that the endophytic bacteria of plants are a subset of the rhizospheric microbial populations (Márquez-Santacruz et al., 2010).
The bacterial populations that inhabit the rhizosphere of wheat plants have mainly been studied by culture methods (Germida & Siciliano, 2001). However, it is known that only a small percentage of these populations can be grown in laboratory conditions. Therefore, molecular methods can give a better indication of the actual populations of bacteria that live in this environment (Hernández-León et al., 2010). Also, there have recently been several metagenomic projects, either trying to know the microbial diversity or building metagenomic libraries, in order to discover molecules or compounds of interest (Handelsman, 2004; Hernández-León et al., 2010). However, before carrying out a metagenomic project it is essential to know how diverse the environment is and whether it is a good candidate to be studied in a future research.
Thus, in this work we studied the bacterial diversity of the rhizosphere of wheat plants in an agricultural crop by PCR amplification of 16S rDNA genes and sequencing. Homology and phylogenetic analysis suggest that there are different genera that can be associated with biocontrol and PGPR bacteria of wheat plants. Finally, diversity analysis shows that this microenvironment is highly diverse, being a good candidate for other metagenomic studies.
MATERIALS AND METHODS
Sampling and physico-chemical analysis of rhizospheric soil. To study the soil microbial community, samples were taken from the rhizosphere of wheat plants near the city of Zamora Michoacan (19° 59' N, 102° 17' W, 1560 m.a.s.l.). Ten plants with their roots and rhizospheric soil were collected in the month of January, when the plants had a month from being planted. Rhizospheric soil samples were taken at 10 cm depth and transported on ice to be stored at 4° C for immediate analysis in the Lab. The rhizosphere was separated from the root and stored at -4° C.
The soil was a clay loam type, with 23.48% sand, 38.57% clay and 38% silt, and a pH of 7.17. Fertilization contained 3.73% organic matter, 16.9 ppm inorganic nitrogen, 653 ppm potassium, 4938 ppm calcium, 1416 ppm magnesium, 153 ppm sodium, 22.8 ppm iron, 0.91 ppm zinc, 55.9 ppm manganese and 2.55 ppm copper. These physico-chemical conditions resulted optimal for wheat crop.
DNA extraction. Total DNA extraction from rhizosheric soil samples were done by using the MO-BIO PowerSoil® DNA Isolation Kit, following manufacturer instructions. Extracted DNA solution was completely transparent and was checked by gel electrophoresis and stained with ethidium bromide (4 µg/mL).
Polymerase chain reaction amplification of the bacterial 16S rRNA genes and contruction of the library. Ribosomal 16S rDNA genes were amplified using the universal bacterial primers Fd1, 5'-CAGAGTTTGATCCTGGCTCAG-3' (forward) and Rd1, 5'-AAGGAGGTGATCCAGCC-3' (reverse), corresponding to positions 8 to 28 and 1526 to 1542 from the Escherichia coli 16S rDNA gene, respectively (Weisburg et al., 1991). The following Polymerase Chain Reaction (PCR) conditions were used: an initial denaturation at 95 °C for 3 min; 30 cycles of 1 min at 95 °C for denaturation, 1 min at 53 °C for annealing, and 2 min at 72 °C for extension, and a final extension step at 72 °C for 5 min. Polymerase chain reaction amplifications were performed with a TC-412 Techne Thermal Cycler. GoTaq® Master Mixes tubes (Promega) were used (tubes were supplied with enzyme, magnesium, dNTPs, and buffer). Only 0.1 µg template DNA and 50 pmol of each primer were added to each tube.
PCR products were gel purified from 1% agarose gels by using the Wizard® SV Gel and PCR Clean-Up System (Promega) according to manufacturer instructions. The purified PCR fragments were then cloned into the pGEM-T Easy Vector (Promega), and the resulting ligation products were used to transform competent E. coli cells. Positive white clones were detected on LB medium containing 80 µg/mL X-Gal and 0.5 mM IPTG.
Sequencing and analysis of the 16S rDNA genes. Ninety-six isolated plasmids from E. coli clones were isolated and corroborated for insert cloning, and were commercially sequenced. The possibility to obtain chimeric sequences was analyzed by using the CHIMERA_ CHECK program of the Ribosomal Database Project (Maidak et al., 1999). Sequences considered to be chimeras were discarded, as well as those with no homology to ribosomal genes, resulting in 84 high-quality sequences for analysis. All sequences obtained were compared with sequences in the GenBank (NCBI) database by using the BLASTN program, to obtain the best matching sequences.
Phylogenetic and statistical analysis. A multiple sequence alignment was generated with ClustalX, and the phylogenetic analysis of the 16S DNA sequences was carried out with the MEGA 4.0 program (Tamura et al., 2007). To obtain a confidence value for the aligned sequence dataset, a bootstrap analysis of 1000 replications was done. Phylogenetic trees were constructed by the neighbor-joining method based on Kimura's two-parameter distance (Kimura, 1980). The eubacterial diversity of the 16S rDNA library was evaluated by a Shannon-Wiener test (Krebs, 1985).
RESULTS
Metagenomic DNA isolation and PCR amplification of 16S rDNA genes. The total rhizosphere DNA was isolated and purified for use as a template in PCR reactions (Fig. 1a). Figure 1b shows the single band of approximately 1.5 kb amplified with universal primers FD1 and RD1 (Weisburger et al., 1991), corresponding to the 16S rDNA of eubacteria.

Fig. 1. (a) Total volume of DNA from rhizosphere of wheat plants. (b) Standard DNA marker and PCR amplification of 16S rDNA genes with approximate size of 1.5 kb.
Fig. 1. (a) Volumen total de ADN de la rizósfera de plantas de trigo. (b) Marcador de ADN estándar y amplificación PCR de genes 16S rDNA con un tamaño aproximado de 1,5 kb.
Bacterial diversity associated with rhizosphere of wheat plants. High-quality sequences were obtained from 84 of the 16S rDNA gene clone library from wheat plants rhizosphere. According to the results of homology search, species pertained to the Classes Alfaproteobacteria (7.1%), Betaproteobacteria (15.4%), Gammaproteobateria (36.9%), Deltaproteobacteria (4.7%), Actinobacteria (10.7%), Bacilli (10.7%) and Clostridia (2.3%). Also, some sequences (10.7%) showed high similarity to uncultured bacteria (Fig. 2).
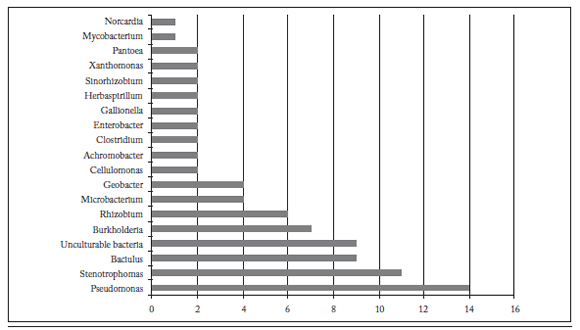
Fig. 2. Bacterial diversity found in the 16S rDNA clone library from the rhizosphere of wheat plants.
Fig. 2. Diversidad bacteriana observada en la biblioteca clonal 16S rADN de la rizósfera de plantas de trigo.
The percentages of identity with matches found in the databases of the National Center for Biotechnology Information (NCBI) and the identified OTUs are presented in Table 1. It also reports the identity of the closest match to the species found in the 16S rDNA gene library from the rhizosphere of wheat plants. The three dominant genera found were Pseudomonas with 14 of the 84 clones (16.6%), followed by Stenotrophomonas with 11 clones (13%) and Bacillus with 9 clones (10.7%). The above mentioned genera comprised more than 40% of the clones analyzed, and they were the most abundant representatives of the bacterial populations from the wheat rhizosphere. We also found 16S ribosomal sequences that showed high identity with non-culturable bacteria (9 clones, 10.7%). Other genera found are some that have been reported previously that are common inhabitants of soil and rhizosphere of plants, such as Burkholderia, Rhizobium, Sinorhizobium and Xanthomonas, as shown in Figure 3.
Table 1. Percentage identity by OTUs with NCBI match of bacterial 16S rDNA sequences from rhizosphere of wheat.
Tabla 1. Identidad porcentual por OTUs con ajuste NCBI de secuencias bacterianas 16S rDNA de la rizósfera de trigo.

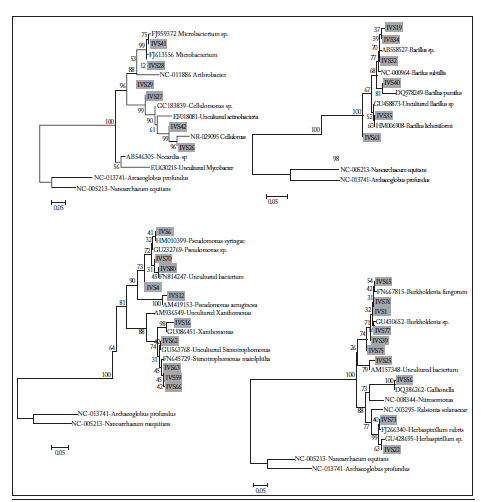
Fig. 3. Dendograms showing the Phylogenetic relationships among the inhabitants of different soil and rhizosphere bacteria of plants. A multiple sequence alignment was generated with ClustalX, and the phylogenetic analysis of the 16S DNA sequences was carried out with the MEGA 4.0 program. To obtain a confidence value for the aligned sequence dataset, a bootstrap analysis of 1000 replications was done. Phylogenetic trees were constructed by the neighbor-joining method based on Kimura's two-parameter distance. Archaeoglobus profundus and Nanoarchaeum equitans were used as outgroups. Clones are indicated in grey squares. See text for details.
Fig. 3. Dendrogramas que muestran las relaciones filogenéticas entre los habitantes de diferentes bacterias del suelo y de la rizósfera de las plantas. Se generó un alineamiento de secuencias múltiples con ClustalX, y el análisis filogenético de las secuencias de 16S ADN se realizó con el programa MEGA 4.0. Se realizó un análisis bootstrap de 1000 repeticiones para obtener un valor de confianza para la serie de datos de secuencia alineada. Se construyeron árboles filogenéticos por el método de unión de vecinos basado en la distancia entre dos parámetros de Kimura. Archaeoglobus profundus y Nanoarchaeum equitans se usaron como grupos exteriores. Los clones se indican con cuadrados grises. Ver el texto para detalles.
Phylogenetic and richness analysis. Some sequences were used for phylogenetic analysis. The most representative are shown in Figure 4, which generally are grouped with clades that belong to bacteria with a ubiquitous distribution and common inhabitants of bulk-soils and rhizosphere environments of different plants. Regarding the diversity analysis by the Shannon-Wiener test, it resulted in a rate of 3.8 bits per individual, suggesting that the rhizosphere of the study wheat plants is highly diverse.
DISCUSSION
The rhizosphere is defined as the portion of soil that is influenced by plant roots. In this microecosystem the root exudates are rich in nutrients, thus attracting a wide variety of microorganisms, including bacteria that can occupy those spaces, obtain nutrients to grow and limit the growth of pathogens (Weller, 1998). Such is the case of rhizospheric bacteria of the genera Bacillus and Pseudomonas, which are inhabitants of the bulk-soil and rhizosphere (Tomashow, 1996); this paper reports its presence as two of the most abundant in the 16S clone library from the rhizosphere of wheat plants. Studies have shown that different strains of Bacillus, through direct or indirect mechanisms, inhibit or control potential pathogens (Leclère et al., 2005; Chen et al., 2009; León et al., 2009). It has been reported that diverse Bacillus species produce antibiotics with antifungal activity, including lipopeptides (Koumoutsi et al., 2004; Ongena & Jacques, 2008). Lipopeptides such as fengycin, surfactin and members of the iturin family have been studied in detail, and have shown to be effective in suppressing different phytopathogenic organisms, including fungi, bacteria and nemetodes, among others (Ongena & Jacques, 2008; Chen et al., 2009). On the other hand, it has also been suggested that some species of Bacillus may be plant-growth promoting through phytostimulation and volatile production (Idris et al., 2004; Ryu et al., 2004; Velázquez-Becerra et al., 2011). In this sense, bacteria of the genus Pseudomonas, like Bacillus, also have the ability to be PGPR, in addition to show biocontrol activities (Compant et al., 2005; Haas & Defago, 2005). Pseudomonas have rapid growth and are therefore good colonizers in soil. Pseudomonas can use various substrates as nutrients and survive under different stressing conditions. Also its ability to produce various compounds, such as antibiotics, polysaccharides and siderophores are crucial to its success (Weller, 2007). In recent works, Valencia-Cantero et al. (2005) and Santoyo et al. (2010) showed that the strain P. fluorescens ZUM80 can restrict growth of plant pathogenic fungi, such as Fusarium oxysporum, Colletotrichum lindemuthianum, Colletotrichum gloeosporioides and Phytophthora cinnamomi, through the synthesis of iron chelators and other potential antibiotics. Another interesting mechanism of indirect biocontrol employed by Bacillus and Pseudomonas is the ability to Induce Systemic Resistance (ISR) in plants, thus protecting them from diverse phytopathogen infections (Compant et al., 2005; Ongena et al., 2007).
Another genus found in the 16S rDNA library from the wheat plant rhizosphere was Stenotrophomonas. This genus also contains rhizospheric habitants, playing an interesting ecological role on plant-growth promoting activities (Valencia-Cantero et al., 2007).
Other works on studies of diversity in rhizosphere of wheat plants have also shown a wide variety of bacteria (Germida & Siciliano, 2001). Importantly, they analyzed the diversity by using the fatty acid methyl esterified method, but not involving isolation of DNA sequences. These authors reported that the bacteria found belonged to the Gammaproteobacteria and Bacilli Classes, and the predominant genera were Pseudomonas, Arthrobacter, Bacillus, Flavobacterium, Micrococcus, Xanthomonas, Agrobacterium and Enterobacter, among others. This is in agreement with our results, where phylogenetic analysis showed that some representative 16S genes were grouped in clusters with sequences of species that are inhabitants of the rhizosphere of plants.
Cellulomonas, Microbacterium and Geobacter are other genera found in the 16S rDNA clone library. These genera have been found as endophytes in different plants, such as rice, soybean, sweet potato, grapevine, Mexican husk tomato, and maize, playing important ecological roles (Ryan et al, 2008; Marquez- Santacruz et al., 2010). Other genera such as Xanthomonas and Pantoea are also endophytes of plants; however, some species are plant pathogens (Bashan et al., 1982). It means that potential phytopatogenic microorganisms are also resident of the rhizosphere of wheat crops. Another major portion of the library was the presence of ribosomal sequences belonging to non-culturable bacteria. This suggests that even though the vast majority of sequences belonged to rhizospheric bacteria, there are still a significant proportion of organisms that could not be detected by culture methods. Finally, the analysis of the Shannon-Wiener diversity of the 16S rDNA library from the rhizosphere of wheat plants suggests that this environment is highly diverse. This is a prerequisite for other metagenomic studies to discover genes of interest with some application in agricultural biotechnology.
ACKNOWLEDGMENTS
We thank Consejo Nacional de Ciencia y Tecnología, México (Proyecto No. 169346) and Coordinación de la Investigación Científica-Universidad Michoacana de San Nicolás de Hidalgo (2011-2012) for financial support to this project. We also thank M. Govindappa for discussion of results.
REFERENCES
1. Adesemoye, A.O. & J.W. Kloepper (2009). Plant-microbes interactions in enhanced fertilizer-use efficiency. Applied Microbiology and Biotechnology 85: 1-12. [ Links ]
2. Adesemoye, A.O., H.A. Torbert & J.W. Kloepper (2009). Plant growth-promoting rhizobacteria allow reduced application rates of chemical fertilizers. Microbial Ecology 58: 921-929. [ Links ]
3. Ahmad, F., I. Ahmad & M.S. Khan (2008). Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiol Res.163: 173-181. [ Links ]
4. Bashan, Y. (1982). Survival of Xanthomonas campestris pv. vesicatoria in pepper seeds and roots in symptomless and dry leaves in non-host plants and in the soil. Plant Soil 68: 161-170. [ Links ]
5. Chen, X.H., A. Koumoutsi, R. Scholz, K. Schneider, J. Vater, R. Süssmuth, J. Piel & R.J. Borriss (2009). Genome analysis of Bacillus amyloliquefaciens FZB42 reveals its potential for biocontrol of plant pathogens. Biotechnology 140: 27-37. [ Links ]
6. Compant, S., B. Duffy, J. Nowak, C. Clément & E.A. Barka (2005). Use of plant growth-promoting bacteria for biocontrol of plant diseases: principles, mechanisms of action, and future prospects. Applied and Environmental Microbiology 71: 4951-4959. [ Links ]
7. Fauvart, M., A.Sánchez-Rodríguez, S. Beullens, K. Marchal & J. Michiels (2011). Genome sequence of Rhizobium etli CNPAF512, a nitrogen-fixing symbiont isolated from bean root nodules in Brazil. Journal of Bacteriology 193: 3158-3159. [ Links ]
8. Germida, J.J. & S.D. Siciliano (2001). Taxonomic diversity of bacteria associated with the roots of modern, recent and ancient wheat cultivars. Biology and Fertility of Soils 33: 410-415. [ Links ]
9. Haas, D. & G. Défago (2005). Biological control of soil-borne pathogens by fluorescent pseudomonads. Nature Reviews Microbiology 3: 307-319. [ Links ]
10. Handelsman, J. (2004). Metagenomics: application of genomics to uncultured microorganisms. Microbiology and Molecular Biology Reviews 68: 669-685. [ Links ]
11. Hayat, R., S. Ali, U. Amara, R. Khalid & I. Ahmed (2010). Soil beneficial bacteria and their role in plant growth promotion: a review. Annals of Microbiology 60: 579-598. [ Links ]
12. He, Z.L., X.E. Yang & P.J. Stoffella (2005). Trace elements in agroecosystems and impacts on the environment. Journal of Trace Elements in Medicine and Biology 19: 125-40. [ Links ]
13. Idris, E.E.S., O. Makarewicz, A. Farouk, K. Rosner, R. Greiner, H. Bochow, T. Richter & R. Borriss (2004). Extracellular phytase activity of Bacillus amyloliquefaciens FZB 45 contributes to its plant-growth-promoting effect. Microbiology 148: 2097-2109. [ Links ]
14. Kimura, M. (1980). A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution 16: 111-120. [ Links ]
15. Krebs, C.J. (1985). Species diversity. In: C. J. Krebs (ed.), Ecology: the experimental analysis of distribution and abundance, 3th ed. Harper and Row, New York, N.Y. Pp. 507-534. [ Links ]
16. Koumoutsi, A., X.H. Chen, A. Henne, H. Liesegang, G. Hitzeroth, P. Franke, J. Vater & R. Borriss (2004). Structural and functional characterization of gene clusters directing nonribosomal synthesis of bioactive cyclic lipopeptides in Bacillus amyloliquefaciens strain FZB42. Journal of Bacteriology 186: 1084-1096. [ Links ]
17. Leclère, V., M. Béchet, A. Adam, J.S. Guez, B. Wathelet, M. Ongena, P. Thonart, F. Gancel, M. Chollet-Imbert & P. Jacques (2005). Mycosubtilin overproduction by Bacillus subtilis BBG100 enhances the organism's antagonistic and biocontrol activities. Applied and Environmental Microbiology 71: 4577-4584. [ Links ]
18. León, M., P.M. Yaryura, M.S. Montecchia, A.I. Hernández, O.S. Correa, N.L. Pucheu, N.L. Kerber & A.F. García (2009). Antifungal activity of selected indigenous Pseudomonas and Bacillus from the soybean rhizosphere. International Journal of Microbiology ID 572049, 9 pages. doi: 10.1155/2009/572049. [ Links ]
19. Maidak, B.L. et al., (1999). A new version of the RDP (Ribosomal Database Project). Nucleic Acids Research 27: 171-173. [ Links ]
20. Marquez-Santacruz, H.A., R. Hernandez-Leon, M.C. Orozco- Mosqueda, I. Velazquez-Sepulveda, & G. Santoyo (2010). Diversity of bacterial endophytes in roots of Mexican husk tomato plants (Physalis ixocarpa) and their detection in the rhizosphere. Genetics and Molecular Research 9: 2372-80. [ Links ]
21. Ongena, M. & P. Jacques (2008). Bacillus lipopeptides: versatile weapons for plant disease biocontrol. Trends in Microbiology 16: 115-125. [ Links ]
22. Ongena, M., E. Jourdan, A. Adam, M. Paquot, A. Brans, B. Joris, J.L. Arpigny & T. Thornart (2007). Surfactin and fengycin lipopeptides of Bacillus subtilis as elicitors of induced systemic resistance in plants. Environmental Microbiology 9: 1084-1090. [ Links ]
23. Ryan, R.P., K. Germaine, A. Franks, D.J. Ryan & D.N. Dowling (2008). Bacterial endophytes: recent developments and applications. FEMS Microbiology Letters 278: 1-9. [ Links ]
24. Ryu, C.M., J.F. Murphy, K.S. Mysore & J.W. Kloepper (2004). Plant growth-promoting rhizobacteria systemically protect Arabidopsis thaliana against Cucumber mosaic virus by a salicylic acid and NPR1-independent and jasmonic acid-dependent signaling pathway. Plant Journal 39: 381-92. [ Links ]
25. Santoyo, G., E. Valencia-Cantero, Ma. Del C. Orozco-Mosqueda, J.J. Peña-Cabriales & R. Farías-Rodríguez (2010). Role of Siderophores in antagonic activity of Pseudomonas fluorescens ZUM80 against plant fungi. Terra Latinoamericana 28: 53-60. [ Links ]
26. Schroth, M.N. & J.W. Kloepper (1978). Plant growth promoting rhizobacteria on radish. In: Proceedings of the Fourth Conference Plant Pathogenic Bacteria. Ed Station de Pathogenic Vegetable ET (Phytobacteriologic) INRA Angers. Pp 876-882. [ Links ]
27. Tamura, K., J. Dudley, M. Nei & S. Kumar (2007). MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution 24: 1596-1599. [ Links ]
28. Thomashow, L.S. (1996). Biological control of plant root pathogens. Current Opinion in Biotecnology 7: 343-347. [ Links ]
29. Velázquez-Becerra, C., L. Macías-Rodríguez, J. López-Bucio, J. Altamirano-Hernández, I. Flores-Cortez & E. Valencia-Cantero (2011). A volatile organic compound analysis from Arthrobacter agilis identifies dimethylhexadecylamine, an amino-containing lipid modulating bacterial growth and Medicago sativa morphogenesis in vitro. Plant and Soil 339: 329-340. [ Links ]
30. Valencia-Cantero, E., J. Villegas-Moreno, J.M. Sánchez-Yáñez, J.J. Peña-Cabriales & R. Farías-Rodríguez (2005). Inhibición del crecimiento de Fusarium oxysporum por cepas mutantes de Pseudomonas fluorescens Zum80 incapaces de producir sideróforos. Terra Latinoamericana 23: 65-72. [ Links ]
31. Valencia-Cantero, E, E. Hernandez-Calderón, C. Velázquez-Becerra, J.E. López-Meza, R. Alfaro-Cuevas & Y.R. Lopez-Bucio (2007). Role of dissimilatory fermentative iron-reducing bacteria in Fe uptake by common bean (Phaseolus vulgaris L.) plants grown in alkaline soil. Plant and Soil 291: 263-273. [ Links ]
32. Weisburger, W., S.M. Barns, D.A. Pelletier & D.J. Lane (1991). 16S Ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology 173: 697-703. [ Links ]
33. Weller, D.M. (1988). Biological control of soilborne plant pathogens in the rhizosphere with bacteria. Annual Review of Phytopathology 26: 379-407. [ Links ]
34. Weller, D.M. (1998). Biological control of soil borne plant pathogens in the rhizosphere with bacteria. Annual Review of Phytopathology 26: 379-407. [ Links ]
35. Weller, D.M. (2007). Pseudomonas biocontrol agents of soil-borne pathogens: Looking back over 30 years. Phytopathology. 97: 250- 256. [ Links ]














