Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO
Links relacionados
-
 Similares em
SciELO
Similares em
SciELO
Compartilhar
Acta Odontológica Latinoamericana
versão On-line ISSN 1852-4834
Acta odontol. latinoam. vol.29 no.3 Buenos Aires dez. 2016
ARTÍCULOS ORIGINALES
Genetic-relatedness of peri-implants and buccal Candida albicans isolates determined by RAPD-PCR
Adriana M. Bertone1, Alcira C. Rosa2, Natalia Nastri2, Hector D. Santillán3, Yamila Ariza1, Cristina A. Iovannitti1, Virginia M. Jewtuchowicz1,3
1 Department of Microbiology, Parasitology and Immunology, School of Medicine, University of Buenos Aires, Argentina.
2 Department of Microbiology and Parasitology, School of Dentistry, University of Buenos Aires, Argentina.
3 Hospital HIGA Gandulfo, Ministry of Health, Buenos Aires, Argentina.
CORRESPONDENCE Dra. Virginia Marta Jewtuchowicz. Loria 574 18a, Lomas de Zamora, Bs As, CP: 1832. Argentina email: minivirjg@gmail.com
ABSTRACT
Molecular techniques have been used in recent studies to identify a wide range of potential bacterial pathogens in periimplant pockets of the oral cavity. However, the prevalence and molecular epidemiology of yeasts and species distribution related to periimplantitis are as yet unknown. The aim of this study was to determine the prevalence and distribution of yeasts in periimplant biofilm and to study genetic relatedness of Candida albicans. Yeasts recovered from periimplant biofilm samples (n=89) and buccal samples (n=120) were studied in 40 immunocompetent nonsmoking patients who visited the dental clinic of the Asociación Implantodontológica Argentina, Buenos Aires, Argentina, and had received oral rehabilitation with implants for more than five years. Yeasts recovered from samples were studied by typing assays using RAPDPCR. The prevalence of yeasts in the periimplant sulcus was 73% (n=29). C. albicans was the most prevalent species identified in this study population. The RAPD analysis showed identical genotypes in most C. albicans spp. from the two different sampling sites: buccal and periimplant. These findings suggest that periimplant biofilm is an ecological niche that favors the growth of yeast species. Most C. albicans found in periimplant biofilm originate from the endogenous infection caused by commensal strains.
Key words: Implants; Biofilm; Candida albicans; RAPDPCR; Periimplantitis.
RESUMEN
Relación genética de aislamientos de Candida albicans por RAPD-PCR en surcos peri-implantarios de cavidad bucal
Las técnicas moleculares se han utilizado en estudios recientes para identificar una gran diversidad de patógenos bacterianos de surcos periimplantarios de cavidad bucal. Sin embargo, la prevalencia y epidemiología molecular de especies de levaduras en relación con la periimplantitis son aún desconocidas. El objetivo de este estudio fue determinar la prevalencia y distribución de las levaduras en la biopelícula periimplantaria y estudiar la relación genética de Candida albicans. Se estudiaron 40 pacientes inmunocompetentes no fumadores que se asistieron en la clínica dental de la Asociación Implantodontológica Argentina, Buenos Aires, Argentina, y que habían recibido rehabilitación oral con implantes durante más de cinco años. Las levaduras aisladas de las muestras de biopelícula periimplantaria (n = 89) y bucales (n = 120), fueron identificadas por métodos micológicos tradicionales y moleculares. Se obtuvo el ADN de C. albicans y se realizaron estudios moleculares por RAPD PCR. La prevalencia de levaduras en el surco alrededor del implante fue de 73 % (n = 29). C. albicans fue la especie más frecuente identificada en esta población de estudio. El análisis RAPD permitió identificar idénticos genotipos de C. albicans en ambos nichos ecológicos estudiados, periimplantar y bucal. Según los resultados obtenidos, el surco periiplantario es un nicho ecológico que favorece el crecimiento de especies de levaduras del género Candida. La mayoría de los aislamientos de C. albicans periimplantarios se originan a partir de la infección endógena causada por cepas comensales.
Palabras clave: Implantes; Biopelícula; Candida albicans; RAPDPCR; Periimplantitis.
INTRODUCTION
The use of osseointegrated implants, as well as their complications and problems, have increased in recent decades. Successfully osseointegrated titanium implants usually harbor low quantities of plaque and present little marginal inflammation. Supraand subgingival microbiota at well maintained implant sites seem to resemble the microbiota associated with healthy gingiva. An increased proportion of putative periodontal pathogens has been documented at implant sites, suggesting that the periodontal pocket may serve as a reservoir for colonization of titanium implants. Periimplantitis is a chronic progressive marginal infection, defined as an inflammatory reaction that affects the tissuesurrounding osseointegrated dental implants, resulting in the loss of the supporting bone. Microbiota resembling that of adult peiodontitis has been found in periimplantitis1-4. .
Extensive antibiotic treatment and irrigation with chlorhexidine may cause etiological changes. Microorganisms not primarily associated with periodontitis, such as Staphylococcus spp., enterics and Candida spp., have also been isolated 2-5. Molecular techniques have been used in recent studies to identify a wide range of potential bacterial pathogens in periimplant pockets 6,7. However, the prevalence of yeasts and species distribution related to periimplantitis are as yet unknown. The same has been found to be true for dental biofilm 2,8 . Dahlen et al. 9, and Reynaud et al.10 claim that there was colonization by the genus Candida spp. in periodontal pockets, refractory periodontitis3,10,11, and implant failure. Other studies report presence of Candida albicans in the subgingival plaque microbiota of human immuno deficiency virus (HIV) positive individuals 12.
In recent years, several molecular typing methods have been used to characterize Candida spp. isolates and to delineate strain relatedness, the most widely used being polymerase chain reaction (PCR) based methods. Among these, the random amplified polymorphic DNA (RAPD) method of DNA fingerprinting has become quite popular for all infectious fungi and has been successfully applied to assess the genetic relatedness of Candida spp. 13-18. These methods have greatly enhanced knowledge on the epidemiology of oral and subgingival Candida spp., and can provide valuable information through their ability to distinguish distinct isolates of the same species. Some studies have demonstrated that commensal yeasts dominate in oral candidiasis, whereas controversial evidence shows that genetically homogeneous, hypervirulent strains of C. albicans are involved in the disease19. Since there is no available data on the epidemiology of yeasts and genetic characterization of periimplant C. albicans, the aim of this study was to characterize periimplant biofilm and mucosal C. albicans isolates recovered from immuno competent subjects with more than 5 years of implant treatment, and to assay the genetic similarity of C. albicans isolates from the two niches in the same patient by RAPD.
MATERIAL AND METHODS
Study population
This study was approved by the Ethics Committee of the School of Pharmacy and Biochemistry, University of Buenos Aires (Res. 41, File 727.071/10). Yeasts recovered from periimplant plaque (n=89) and buccal samples (n=120) were studied in 40 immunocompetent nonsmoking patients with more than five years of implant treatment on oral prosthesis who attended the dental clinic of the Asociacion Implantodontologica Argentina, Buenos Aires, Argentina. Evaluations included clinical examination and radiographs with clinical measurements: pocket depth (PD), considered regular up to 3 mm around implants, plaque index, gingival index 11,20 and bleeding on probing. Measurements were taken at four sites per tooth (mesial, buccal, distal and lingual positions) on 15 teeth, excluding third molars. Bone resorption was assessed by comparing the radiographic examination in the patients' medical records taken at the time of implant placement to those taken at the appointment for this study. In order to analyze bone resorption, implants were classified into two groups according to time of implant placement: “immediately loaded implants” if they were placed during the same session as tooth extraction or “delayed loaded implants” if they were placed on healed bone, months or years after extraction.
Participation in our survey was voluntary and all patients provided written informed consent. The volunteers were requested to rinse their mouths thoroughly with sterile distilled water, after which sterile swabs were used to take samples from tongue, palate and cheek. The dental professional then isolated the area using cotton rolls and a highspeed suction device. Following removal of the supragingival plaque using a Teflon curette to avoid salivary contamination, periimplant biofilm was collected from the interdental plate by inserting 34 sterile paper points number 303540 for 1530 minutes in the four sites: mesial, buccal, distal and lingual positions. Samples were cultured in a differential chromogenic medium (CHROMagar Candida, Paris, France). Yeast isolates were identified using conventional mycological methods: colony color on the chromogenic medium, micromorphology in agar milk with 1% Tween80 21, carbohydrate assimilation tests using a commercially available kit API ID 32D (BioMerieux, Lyon, France), and specific PCR22.
Random amplified polymorphic DNA (RAPD) analysis
Yeast DNA was isolated using a technique described previously 2224. Five different primers were included in the typing assays. Primer sequences were as follows:
OPA 02 (TGCCGAGCTG), OPA 09 (GGGTAACGCC), M13F (CGACGTTGTAAAACGACGCCCAGT), M13R (CAGGAAACAGCTATGAC), and OCP 5 (GATGACCGCC). They were all used in RAPDPCR, following the method developed by Williams et al.23. Arbitrary amplification was performed in a total volume of 50 μl containing: 1_buffer 2.5 mM MgCl2, 0.2 mM each of the dNTP, 0.5 mM of the primer, 1.25 U Taq DNA polymerase (Invitrogen), and 75 ng of template DNA. The cycling program consisted of 4 min at 94oC, 35 1minute cycles at 94oC, 1 min at 25oC, 2 min at 72oC followed by a final extension of 5 min at 72oC.
These steps were carried out in a Minicycler DNA thermal cycler (TM MJ Research Inc., NY, USA). Products were separated by electrophoresis in 1.4% agarose gel and stained with ethidium bromide. They were visualized under UV light and digitalized by image analyzer software (EPIChemi Darkroom. UVP Laboratory Products, California, USA). Band profiles were analyzed and compared visually. Each band was scored as positive or negative for all isolates; and the presence or absence of each band was recorded for each isolate. The resulting matrix was interpreted using the Treecon program, where isolates were grouped according to the resemblance of their patterns. Based on matrix of similarity coefficients (SC), a dendrogram was generated by the unweighted pair group method using arithmetic averages (UPGMA). The criterion used for genotyping was as follows: arbitrary threshold at an SC of 90% for closely related isolates.
Statistical analysis
Statistical analysis was performed using Statistix 7.0 and the SPSS 11.0 software. Confidence interval was 95% (CI 95%). Fisher and ANOVA were calculated at 95% using the EpiInfo 6.04 program (Atlanta University, GA).
RESULTS
Clinical features
The 40 subjects included in the study ranged in age from 33 to 76 years (mean age 56 years), 50% were female (20/40). None of them had received antibacterial or antifungal agents before this treatment. Of the total population, 68% were nonsmokers. This population had an average of 12.80 teeth and 2.58 implants; 1.85 loaded implants and 0.38 nonloaded implants. Of the total number of original implants (n=103) in the study population, we found that only 89 were present. The percentage of bone resorption in immediately loaded implants (n=13), was significantly higher (p<0.001) than in delayed loaded implants (n=76) (Fig.1).

Fig. 1: Percentage of bone resorption in immediatelyloaded and delayedloaded implants. (N=89).
Comparison of bone resorption in relation to the kind of prosthesis placed on the implants (n=89) showed significantly higher resorption rates (p< 0.001) in the group with removable prostheses (36/48) than in the group with fixed prostheses (6/26) and without load (7/15) (Table 1). Pocket depth (PD) was more than 3mm in 18/40 patients and less than 3 mm in 22/40 patients (Table 2).
Table 1: Study of bone resorption in 89 implants.

Table 2: Pocket depth greater and smaller than 3mm.
Carriage of C. albicans and other yeast species
The prevalence of yeasts in the periimplant sulcus was 73% (n=29, CI 95%:55.984.9). In buccal mucosa, the distribution of yeasts was: 73% in palate and cheek (n=29), CI 95%; 0.559 0.859), and 85% in lingual mucosa (n=34, CI 95%; 70.294.3), representing a high statistically significant prevalence (p<0.001) (Table 3).
Table 3: Prevalence of yeasts in the peri-implant sulcus and mucosa.

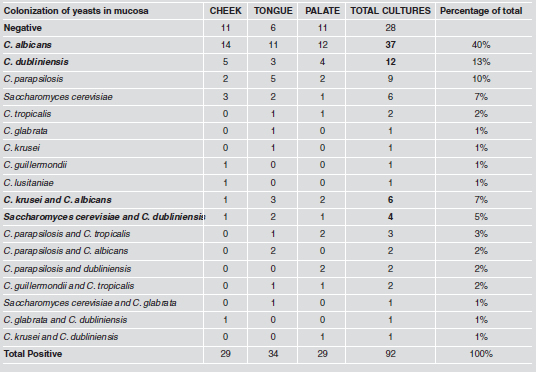
Table 4 summarizes species distribution of yeast isolates in periimplant biofilm and buccal mucosa. Of the 140 yeasts recovered, C. albicans was the species most frequently found in all niches, periimplant and mucosa. The prevalence of C. albicans was 55% (n=22) in periimplant biofilm. Other nonC. albicans spp. and other yeasts were found: C. dubliniensis (n=11), C. parapsilosis (n=5), Saccharomyces cereviciae (n=5), C. krusei (n=2), C. tropicalis (n=1), C. lusitaniae (n=1) and Rhodotorula spp. (n=1). The occurrence of two or three coisolated species was observed in 22/120 buccal mucosa samples. C. albicans and C. krusei (n=6) followed by Saccharomyces cereviciae and C. dubliniensis (n=4) were the associations most frequently observed. The combinations in periimplant sulcus was 16.7% (n=8). Of the associations of the species found, the most predominant were C. dubliniensis with C. krusei, and C. albicans with C. glabrata (2% each) (Table 5).
Table 4: Prevalence of Candida albicans in peri-implant sulcus.
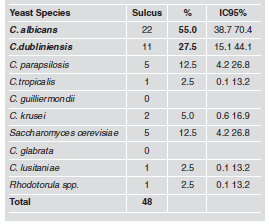
Table 5: Distribution of yeasts in mucosa.
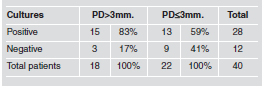
In relation to pocket depth and presence of yeasts, patients with periimplant sulcus >3 mm exhibited an increase in positive cultures (83%, 15/18) compared to negative cultures (17%, 3/18), whereas patients with periimplant sulcus ≤3 mm, positive cultures (59%, 13/22) and negative cultures (41%, 9/22) exhibited much lower discrepancy. This difference was not statistically significant (Table 6).
Table 6: Presence of yeasts in relation to pocket depth.

Of the 89 implants studied, 43 showed no colonization by Candida, of which 23 had bone resorption (53 %) and 20 did not (47%). Of the 46 implants where there was colonization by Candida, 26 had resorption (47%) while the other 20 did not (43%). In all four cases, the percentages were similar. According to these results, periimplant Candida colonization would not be the determining cause of bone resorption around implants. (Fig. 2)

Fig. 2: Percentage of implants with and without Candida colonization.
Implants with removable prostheses exhibited significantly higher (p<0.001) rates of Candida spp. colonization (19/22) than those with fixed prostheses (9/18) (Table 7).
Table 7: Colonization of Candida spp. in implants with removable and fixed prosthesis.

RAPDPCR ASSAY
We selected five RAPD primers, based on their reproducibility, after the prescreening process in order to analyze 68 C. albicans isolates. The number of bands ranged from two to three splitters (M13r) to 12 (M13f). Three of five primers were the most informative (M13f, OPA 9 and OPC5) and generated the highest number of band patterns (10 to 12). The dendrogram generated by the UPGMA clustering method, using the RAPDPCR technique for C. albicans in oral cavity, tongue (LE), palate (PA), cheek (CA), and periimplant sulcus (I) shows similarity coefficient (SC) ranging from 60% to 100%. Thirteen genetic clusters and nine main genotypes were obtained at a similarity coefficient (SC) of 90%, genotypes I, II, III, IV, and V. (Fig 3)
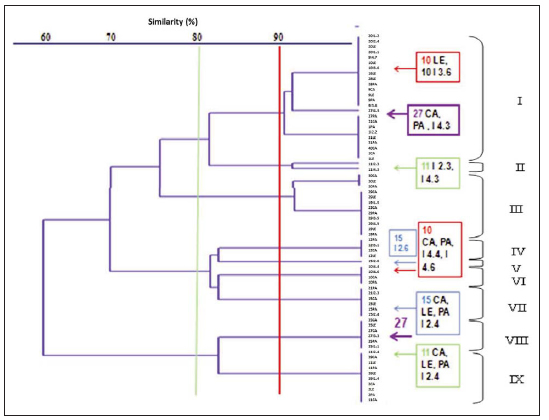
Fig. 3: The dendrogram generated by the UPGMA clustering method, using the coefficient of similarity between RAPDPCR of C. albicans in oral cavity, tongue (LE), palate (PA), cheek (CA), and periimplant sulcus (I) shows that the similarity coefficient (SC) ranged from 60 to 100%. Thirteen genetic clusters and nine main genotypes were obtained at a similarity coefficient (SC) of 90%, genotypes I, II, III, IV and V.
DISCUSSION
In this study, 40 immunocompetent adult patients with more than 5 years' treatment were recruited and grouped according to their health status and pocket depth into periimplantitis or healthy. As expected, patients with periimplantitis presented more infectious sites, including higher rates of percentage similarity (PS) (Anova Test p< 0.001). Eightynine periimplant sulcus samples and 120 swabs from buccal mucosa were cultured directly in CHROMagar Candida medium to enable the presumptive identification of C. albicans or C. dubliniensis, C. tropicalis and C. krusei. This also enabled identification of the presence of infections caused by more than one species simultaneously. Similar findings have been reported by other authors who analyzed other populations22,25,26-28. The prevalence of yeasts in sulcus was 73% (n=29), showing that the surrounding ecological niche and periimplant sulcus enabled yeast growth. Other studies have reported the presence of Candida spp. in periimplant lesions29,30, and found Candida spp. in 55% of periimplant sites. The comparison of yeast distribution in relation to clinical markers of periimplantitis revealed no significant difference in the prevalence of yeasts at sites with PD >3 mm or at sites with bone resorption.
These findings revealed the presence of yeast species in periimplant sulcus as well at sites with or without periimplantitis. Of the 120 buccal mucosa samples studied here, the tongue was the site with highest prevalence of Candida spp. (85, CI95%, 0.702 0.943), in contrast to cheek and palate, with a statistically significant difference (p<0.001).
Candida spp. prevalence was higher in our study than in previously reported series31-34 in which it ranged from 25% to 65%, suggesting that the presence of implants in our study population increases prevalence. In relation to the type of implant rehabilitation −fixed or removable− the latter yielded significantly higher (p<0.001) prevalence of yeasts. It is worth noting that these findings suggest that periimplant plaque is an ecological niche that favors the growth of yeast species; especially in implants with removable rehabilitation, even though they can be removed for cleaning. Moreover, these implants are made of acrylic, which favors adhesion of Candida spp. These are the first data results reported in Argentina. The use of buccal devices induces alterations within the oral cavity. Hagg et al.35 observed that the presence of prosthesis or other buccal devices increases the number of Candida spp., not only at the site but throughout the mucosa. Dental prostheses are made of acrylic resins in which surface defects favor the development of plaque and prevent its removal36. The surface of the prosthesis is very porous and thus susceptible to being colonized by large numbers of microorganisms, which may give rise to different pathologies in the oral cavity. Comparison of the two study samples showed “high” concordance, with colonization or infection by the same yeast in both ecological niches in 95% of the patients (Kappa=0.8).
In relation to the distribution of yeast species, C. albicans spp. was the most prevalent (55%, n=22), but it is important to highlight that nonC. albicans spp. were also found in periimplant sulcus: C. dubliniensis 27.5% (n=11), C. parapsilosis 12.5% (n=5), Saccaromyces cereviciae 12.5% (n=5), C. tropicalis, C. lusitaniae and Rhodotorula spp. 2.5% (n=1), and C. krusei 5% (n=2), (Table 1). Many of these less prevalent species are emerging and characterized by the presence of diminished sensitivity to antifungals37. No data is available in the literature reviewed.
Epidemiological surveillance is very important for identifying the prevalence of yeast species in the biofilm of periimplant sulcus since they create reservoirs for opportunistic microorganisms which, in certain clinical situations such as patients with immune deficiencies, play a significant role in diseases such as buccal candidiasis and disseminated diseases 34, 38. In this study, C. albicans isolates from the buccal cavity and periimplant sulcus of the same patient were considered to be closely related in 90% of the cases (16/20) according to RAPDPCR. Similarity among isolates from both ecological niches suggests that the source of C. albicans colonization in periimplant biofilm is the patient's buccal cavity. Thus, it can be assumed that most C. albicans spp. found in periimplant biofilm originate from endogenous infection by commensal strains. Coincidentally, other authors have found identical genetic patterns in yeasts from different anatomical sites in the same patient. However, the results obtained highlight the fact that the same patient carries different species39. It is important to consider that C. albicans colonization in periimplant sulcus could also occur due to the presence strains adaptable to the periimplant environment, which is likely as a result of genetic variations such as gene conversion and/or chromosomal translocations 15, 19. To date, scientific literature has not provided any information on the genetic characterization of C. albicans isolates in periimplant sulcus. Hence, yeast isolates were analyzed by RAPDPCR, which has proved to be a rapid, simple, costeffective technique and discriminatory for the molecular typing of C. albicans isolates. Other authors have used the same techniques to assay several yeasts species 13, 15-17, 22. This is the first study conducted in Argentina on the molecular characterization of clinical C. albicans isolates in periimplant sulcus by RAPDPCR. We confirm that the periimplant plaque is an ecological niche that favors the growth of yeast species; especially in implants with removable rehabilitation.
C. albicans spp. were the most prevalent in periimplant samples, but it is important to highlight that nonC. albicans spp. were also found in periimplant sulcus, e.g. C. dubliniensis, C. parapsilosis, Saccaromyces cereviciae, C. tropicalis, C. lusitaniae and C. krusei. The findings suggest that most periimplant C. albicans originate from endogenous infection by commensal strains.
ACKNOWLEDGEMENTS
The authors would like to thank Prof. Ricardo Macchi for his advice on statistics (School of Dentistry, University of Buenos Aires, Argentina). This study was supported by grants UBACYT 20020100200204 and 20020150100219BA from the University of Buenos Aires.
1. Mombelli A. Aging and the periodontal and periimplant microbiota. Periodontol 2000 1998; 16:44-52. [ Links ]
2. Marsh PD. Dental plaque: biological significance of a biofilm and community lifestyle. J Clin Periodontol 2005; 32:7-15. [ Links ]
3. Listgarten MA, Lai CH, Young V. Microbial composition and pattern of antibiotic resistance in subgingival microbial samples from patients with refractory periodontitis. J Clin Periodontol 1993; 64:155-161. [ Links ]
4. Marsh PD. Plaque as a biofilm: pharmacological principles of drug delivery and action in the suband supragingival environment. Oral Dis 2003; 9:16-22. [ Links ]
5. Pye AD, Lockhart DE, Dawson MP, Murray CA, Smith AJ. A review of dental implants and infection. J Hosp Infect. 2009; 72:104-110. [ Links ]
6. Maximo MB, De Mendonca AC, Renata Santos V, Figueiredo LC, Feres M, Duarte PM. Shortterm clinical and microbiological evaluations of periimplant diseases before and after mechanical antiinfective therapies. Clin Oral Implants Res 2009; 20:99-108. [ Links ]
7. Norowski PA Jr, Bumgardner JD. Biomaterial and antibiotic strategies for periimplantitis: a review. J Biomed Mater Res B Appl Biomater 2009; 288:530-543. [ Links ]
8. Jarvensivu A, Hietanen J, Rautemaa R, Sorsa T, Richardson M. Candida yeasts in chronic periodontitis tissues and subgingival microbial biofilms in vivo. Oral Dis 2004; 10:106-112. [ Links ]
9. Dahlen G, Wikstrom M. Occurrence of enteric rods, staphylococci and Candida in subgingival samples. Oral Microbiol Immunol 1995; 10:42-46. [ Links ]
10. Reynaud AH, Nygaard-Ostby B, Boygard GK, Eribe ER, Olsen I, Gjermo P. Yeasts in periodontal pockets. J Clin Periodontol 2001; 28:860-864. [ Links ]
11. Pizzo G, Giammanco GM, Pecorella S, Campisi G, Mammina C, D'Angelo M. Biotypes and randomly amplified polymorphic DNA (RAPD) profiles of subgingival Candida albicans isolates in HIV infection. New Microbiol 2005; 28:75-82. [ Links ]
12. Brady LJ, Walker C, Oxford G, Stewart C, Magnusson I, McArthur W. Oral diseases, mycology and periodontal microbiology of HIV1infected women. Oral Microbiol Immunol 1996; 11:371-380. [ Links ]
13. Jain P, Khan ZK, Bhattacharya E, Ranade SA. Variation in random amplified polymorphic DNA (RAPD) profiles specific to fluconazoleresistant and sensitive strains of Candida albicans. Diag Microbiol Infect Dis 2001; 41:113-119. [ Links ]
14. Dassanayake RS, Samaranayake LP. Amplificationbased nucleic acid scanning techniques to assess genetic polymorphism in Candida. Crit Rev Microbiol 2003; 29:1-24. [ Links ]
15. Montour L, Tey R, Xu J. Isolation of Candida dubliniensis in an aboriginal community in Ontario, Canada. J Clin Microbiol 2003; 41:3423-3426. [ Links ]
16. Samaranayake YH, Samaranayake LP, Dassanayake RS, Yau JY, Tsang WK, Cheung BP, Yeung KW. Genotypic shuffling' of sequential clones of Candida albicans in HIVinfected individuals with and without symptomatic oral candidiasis. J Med Microbiol 2003; 52:349-359. [ Links ]
17. Costa F, Manaia CM, Figueiral MH, Pinto E. Genotypic analysis of Candida albicans isolates obtained from removable prosthesis wearers. Lett Appl Microbiol 2008; 46:445-449. [ Links ]
18. Jewtuchowicz VM, Mujica MT, Malzone MC, Cuesta A, Nastri ML, Iovannitti CA, Rosa AC. Genetic relatedness of subgingival and buccal Candida dubliniensis isolates in immunocompetent subjects assessed by RAPDPCR. J Oral Microbiol 2009; 15:17. DOI: 10.3402/jom.v1i0.2003. [ Links ]
19. Song X, Eribe ER, Sun J, Hansen BF, Olsen I. Genetic relatedness of oral yeasts within and between patients with marginal periodontitis and subjects with oral health. J Periodon Res 2005; 40:446-452. [ Links ]
20. Silness J, Loe H. Periodontal Disease in Pregnancy. Correlation between Oral Hygiene and Periodontal Condition. Acta Odontol Scand 1964; 22:121-135. [ Links ]
21. Jitsurong S, Kiamsiri S, Pattararangrong N. New milk medium for germ tube and chlamydoconidia production by Candida albicans. Mycopathologia 1993; 123:95-98. [ Links ]
22. Jewtuchowicz VM, Mujica MT, Brusca MI, Sordelli N, Malzone MC, Pola SJ, Iovannitti CA, Rosa AC. Phenotypic and genotypic identification of Candida dubliniensis from subgingival sites in immunocompetent subjects in Argentina. Oral Microbiol Immunol 2008; 23:505-509. [ Links ]
23. Scherer S, Stevens DA. Application of DNA typing methods to epidemiology and taxonomy of Candida species. J Clin Microbiol 1987; 25:675-679. [ Links ]
24. Duran EL, Mujica MT, Jewtuchowicz VM, Finquelievich JL, Pinoni MV, Iovannitti CA. Examination of the genetic variability among biofilmforming Candida albicans clinical isolates. Rev Iberoam Micol 1987; 24:268-271. [ Links ]
25. Williams JG, Kubelik AR, Livak KJ, Rafalski JA, Tingey SV. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 1990; 18:6531-6535. [ Links ]
26. Soll DR. The ins and outs of DNA fingerprinting the infectious fungi. Clin Microbiol Rev 2000; 13:332-370. [ Links ]
27. Mujica MT, Finquelievich JL, Jewtuchowicz V, Iovannitti CA. Prevalence of Candida albicans and Candida nonalbicans in clinical samples during 19992001. Rev Argent Microbiol 2004; 36:107-112. [ Links ]
28. Jewtuchowicz VM, Brusca MI, Mujica MT, Gliosca LA, Finquelievich JL, Lovannitti CA, Rosa AC. Subgingival distribution of yeast and their antifungal susceptibility in immunocompetent subjects with and without dental devices. Acta Odontol Latinoam 2007; 20:17-22. [ Links ]
29. Laine P, Salo A, Kontio R, Ylijoki S, Lindqvist C, Suuronen R. Failed dental implants clinical, radiological and bacteriological findings in 17 patients. J Craniofac Surg 2005; 33:212-217. [ Links ]
30. Salvi GE, Furst MM, Lang NP, Persson GR. Oneyear bacterial colonization patterns of Staphylococcus aureus and other bacteria at implants and adjacent teeth. Clin Oral Implants Res 2008; 19:242-248. [ Links ]
31. Kleinegger CL, Lockhart SR, Vargas K, Soll DR. Frequency, intensity, species, and strains of oral Candida vary as a function of host age. J Clin Microbiol 1996; 34:2246-2254. [ Links ]
32. Aguirre Urizar JM. Oral candidiasis. Rev Iberoam Micol 2002; 19:17-21. [ Links ]
33. Negroni M, Gonzalez MI, Levin B, Cuesta A, Iovanniti C. Candida carriage in the oral mucosa of a student population: adhesiveness of the strains and predisposing factors. Rev Argent Microbiol 2002; 34:22-28. [ Links ]
34. Luque AG, Biasoli MS, Tosello ME, Binolfi A, Lupo S, Magaro HM. Oral yeast carriage in HIVinfected and noninfected populations in Rosario, Argentina. Mycoses 2009; 52:53-59. [ Links ]
35. Hagg U, Kaveewatcharanont P, Samaranayake YH, Samaranayake LP. The effect of fixed orthodontic appliances on the oral carriage of Candida species and Enterobacteriaceae. Eur J Orthod 2004; 26:623629. [ Links ]
36. Verran J, Maryan CJ. Retention of Candida albicans on acrylic resin and silicone of different surface topography. J Prosthet Dent 1997; 77:535-539. [ Links ]
37. Hazen KC. New and emerging yeast pathogens. Clin Microbiol Rev 1995; 8:462-478. [ Links ]
38. Gonzalez Gravina H, Gonzalez de Moran E, Zambrano O, Lozano Chourio M, Rodriguez de Valero S, Robertis S, Mesa L. Oral Candidiasis in children and adolescents with cancer. Identification of Candida spp. Med Oral Patol Oral Cir Bucal 2007; 12:419-423. [ Links ]
39. Badoc C, De Meeus T, Bertout S, Odds FC, Mallie M, Bastide JM. Clonality structure in Candida dubliniensis. FEMS Microbiol Lett 2002; 209:249-254. [ Links ]